Envion Air Purifier 640, 300 sq ft Room Capacity, Black - envion 49328
Ok the first drain hurt me like heck again I got up and dance around I had to pinch the tube to stop the pain once but I’m suprised my machince didn’t beep guess I didn’t pinch it that long . Ok drain done then first fill then first dwell one out of three so went through that three the dwell are pretty long the nurse decided to only have ne do a full drain at first because my pain is so bad so she had that set up the other two left some in and at the very end she leaving 500 in me cause of my pain .
I wasn't getting much sleep on the machine, I would wake up with each exchange (I'm a light sleeper I guess), it would never drain properly unless I stood and danced.
Loss of nuclear QKI-5 causes reciprocal changes of NEAT1 isoforms.A, schematic of the putative QKI recognition element (QRE) upstream of the human NEAT1 PAS. B, schematic depicting the deletion of QKI-5 exon 7c by CRISPR-Cas9. Scissors depict cleavage sites by sgRNAs. C, RT-qPCR detection of QKI-5 mRNA in the U373 ΔQKI-5 clone compared to control. Data are shown as mean ± SD from 6 biological replicates, normalized to β-actin, and compared using the ΔΔCT method. Unpaired Student’s t test was used, ∗∗p < 0.01. D, immunoblot analysis of QKI-5 protein levels in the U373 control, the two NEAT1 ΔPAS clones, and the ΔQKI-5 clone. eIF5 was used as a loading control. E–G, RT-qPCR detection of (E) NEAT1_1, (F) NEAT1_2, and (G) quantification of the ratio of NEAT1_2 to NEAT1_1 steady state in U373 ΔQKI-5 clone compared to control. Data are shown as mean ± SD from 3 or 4 biological replicates, normalized to β-Actin, and compared using the ΔΔCT method. Unpaired Student’s t test was used, ∗p < 0.05, ∗∗p < 0.01. NEAT1, nuclear paraspeckle assembly transcript 1; PAS, proximal polyadenylation site; QKI, quaking; RT-qPCR, quantitative RT-PCR; sgRNA, synthetic guide RNA.
This work was supported by National Institutes of Health under the following awards: National Institute of Neurological Disorders and Stroke (R01NS110110 to Y. F., R01NS118819 to Y. F and B. Y., and R01NS100967 to R. D. R.), National Institute on Aging (R01AG078937 to B. Y.), and National Institute of Mental Health (F31MH127915 to P. M. Z.). The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Oh wow, sorry to hear that friday night ask last night was bad. I would be talking to the Surgeon if the dialysis center won't listen to you. Did you ever have a follow up with the Surgeon after your catheter was put in place? Ask the surgeon if he would do x-ray or something to check it out for placement issues. I mean, he's the one who put it there!
The authors thank Dr Archa H. Fox for the feedback and guidance on the RNA-FISH work and for sharing the NONO antibody. We thank Luke A. Knudson for the guidance with our Imaris analysis. We thank Dr Anita H. Corbett for the critical review of the manuscript and intellectual contribution. We also thank Dr M. Lee Cato for the critical review and feedback of the manuscript.
Rothco 1946 Gold Deluxe Concealed Weapons Permit Badge. This product has zinc alloy with gold plate and pin back closure It is a gold color and has ...
Poly(A) RNA was isolated using the NEBNext Poly(A) mRNA Magnetic Isolation Module (NEB, E7490L), following the manufacturer’s instructions.
yeah the trash is overwhelming I had a small trash can in my bathroom and said just isn’t going to work so I bought a big stainless steel can for bathroom . Yes one thing it didn’t say on my cheat sheet is wiping down the transport set with Alcavis but my nurse told me to have out one gauge for night and one for morning so I always wipe it down before I disconnect just like we did on manual . I’m glad your box work and you had minimal drain pad . I’m on day three of tartar bowl drain pain iso I’m praying that continues.
That's okay I got the bottle and it sat for a couple of days before I remembered to put it in the kitchen as a reminder. Not sure if it's the dialysis or what but my concentration/memory is shot. Sometimes I forget to eat, well I'm not hungry a lot of times.
Cato 11001 hreplacement
They really don't like skipping so I'll fill before, make the appt dwelling, and then drain while there. If it runs too late I'll fill while there.
Ok, good to know you have a connection with the Surgeon once the time comes so you can get his opinion and/or get him to look into it. Really glad they will let you use the bypass when needed. NOONE should be going though torture with all this. That's what I was feeling with manuals when i called it and just stopped. So do what you gotta do. Wish I had more ideas for you, but other than doing what you're doing, there's not much you can do. I really hope yours does like mine and stops on it's on for the most part after another week or two. Time will tell. Till then, bypass the crap out of it!
Cato 11001 hdimensions
Being on the cycler at night should make it easy for you to go to your appts during the day, and any your husband needs. I've got my monthly doc visit at 1030 on the 15th and am trying to figure out how to do my first exchange and make the dr appt without getting up at 5 am to fill, dwell, drain. And I still have 3 dogs who have to be taken care of. I get a headache thinking about it.
When do you place your next Baxter order? Did you get it all figured out? They want me to place my order around the 6th of each month for shipment to arrive around the 20th.
The Solution bags come in 6000 and 3000...Have you PD Nurse change the setup so you can use the 3000...so much easier to manipulate ...the box with still be heavy because there will be 4 instead of just 2, but so much easier to pull out a box
All images were obtained using a Nikon Eclipse TE2000 (Nikon) widefield fluorescence microscope with a 60× objective. Z-series were acquired at 0.2 μm steps, and image stacks were deconvolved using AutoQuant X3 software (https://mediacy.com/autoquant-deconvolution/; Media Cybernetics) and a 3-D blind algorithm. All acquisition parameters were kept consistent for all samples. The “Coloc” module of the Imaris software (https://imaris.oxinst.com/; Bitplane) was used for analysis and quantification of paraspeckle number and paraspeckle area. All intensity thresholds were set the same across all samples. Representative images were prepared using the FIJI software package (https://imagej.net/software/fiji/; ImageJ), and dot plots were generated with GraphPad Prism 10.0 (https://www.graphpad.com/; GraphPad Software).
The long noncoding RNA nuclear paraspeckle assembly transcript 1 (NEAT1) is involved in a variety of human cancers. Two overlapping NEAT1 isoforms, NEAT1_1 and NEAT1_2, are produced through mutually exclusive alternative 3′ end formation. Previous studies extensively investigated NEAT1 dysregulation in tumors, but often failed to achieve distinct quantification of the two NEAT1 isoforms. Moreover, molecular mechanisms governing the biogenesis of NEAT1 isoforms and the functional impacts of their dysregulation in tumorigenesis remain poorly understood. In this study, we employed an isoform-specific quantification assay and found differential dysregulation of NEAT1 isoforms in patient-derived glioblastoma multiforme cells. We further showed usage of the NEAT1 proximal polyadenylation site (PAS) is a critical mechanism that controls glioma NEAT1 isoform production. CRISPR-Cas9–mediated PAS deletion reduced NEAT1_1 and reciprocally increased NEAT1_2, which enhanced nuclear paraspeckle formation in human glioma cells. Moreover, the utilization of the NEAT1 PAS is facilitated by the RNA-binding protein quaking (QKI), which binds to the proximal QKI recognition elements. Functionally, we identified transcriptomic changes and altered biological pathways caused by NEAT1 isoform imbalance in glioma cells, including the pathway for the regulation of cell migration. Finally, we demonstrated the forced increase of NEAT1_2 upon NEAT1 PAS deletion is responsible for driving glioma cell migration and promoting the expression of genes implicated in the regulation of cell migration. Together, our studies uncovered a novel mechanism that regulates NEAT1 isoforms and their functional impacts on the glioma transcriptome, which affects pathological pathways of glioma, represented by migration.
Alterations of NEAT1 isoform levels influence paraspeckle abundance.A, representative RNA-FISH images of U373 GBM control and two NEAT1 ΔPAS clones stained for NEAT1_2 (red) and immunofluorescence of NONO (green). Nuclei were stained with DAPI (blue). Arrows indicate representative paraspeckles, identified by colocalization of the NEAT1 FISH probe and NONO IF. Field image scale bar represents 25 μm; zoomed image scale bar represents 10 μm. B and C, dot plot quantification of (B) number of paraspeckles (colocalized foci) and (C) area of individual paraspeckle (μm2) in control and two U373 NEAT1 ΔPAS clones. Data are shown as the mean ± SD of 50 nuclei. Statistical significance was calculated using one-way ANOVA with Dunnett multiple comparison’s test, ns = not significant, ∗∗p < 0.01. D, representative RNA FISH images of U373 GBM control and two NEAT1 ΔPAS clones stained for Total NEAT1 (red) and immunofluorescence of NONO (green). Nuclei were stained with DAPI (blue). Field image scale bar represents 25 μm; zoomed image scale bar represents 10 μm. E and F, dot plot quantification of (E) number of paraspeckles (colocalized foci) and (F) area of individual paraspeckle (μm2) in control and two U373 NEAT1 ΔPAS clones. Arrows indicate paraspeckles. Data are shown as the mean ± SD of 50 nuclei. Statistical significance was calculated using one-way ANOVA with Dunnett multiple comparison’s test, ns = not significant, ∗p < 0.05 and ∗∗∗p < 0.001. DAPI, 4′,6-diamidino-2-phenylindole; GBM, glioblastoma multiforme; IF, immunofluorescence; NEAT1, nuclear paraspeckle assembly transcript 1; PAS, proximal polyadenylation site.
Increasing the solution % only pulls out more water (i.e. more UF), it doesn't pull out more toxins. They have to either add more exchange or increase the volume to meet the adequacy.
I am a very light sleeper I don’t think the Machince is so loud I hear a little noise it more like a wave sound. I will see if I can record it tonight and if somehow I can put a video on here. I thought it would be louder then it was .
Guess I'll have to have my husband help me then. I finally got rid of the 16 (8 yellow, 8 green) boxes I got as a duplicate order. I'm taking them to the center today along with my appt with the neph. I have some questions for him.
Yes it will...but I like to check my drain bag each day for Fibrin...blood...or cloudiness...I can't really see that if I use the toilet...My PD Nurse said if I see blood or cloudiness that I will have to take it to Clinic for her to look at it...If I find Fibrin then I inject Heparine into my bag the next night
I just dance the end of the drain cause that when pain hit☹️yeah we could be dancing for something fun not dialysis lol . I’m a light sleeper too if this drain pain don’t end I may be following suit going to manual but I promise pd nurse and doctor I will give it more time. But I rather be a dancing Queen somewhere else🤪
Well the first night went well. A lot of "did I shut off air", making sure I was hands, looking for best set up. I did get a 5 gallon bucket to drain into and holy cow it was almost full.
Reciprocal changes of NEAT1 isoforms broadly altered the GBM transcriptome.A, volcano plot of one NEAT1 ΔPAS clone indicates differentially expressed genes (DEGs) in U373 cells due to loss of the NEAT1 PAS. Blue dots represent DEGs with significant reductions, whereas red dots represent DEGs with a significant increase. Black dots represent genes that do not change significantly (FDR > 0.05) upon loss of the NEAT1 PAS. B, bar chart indicating the number of identified upregulated and downregulated DEGs in the two NEAT1 ΔPAS clones, respectively. C, Venn diagram shows significant overlap of increased and decreased DEGs between the two NEAT1 ΔPAS clones. D, bar plot indicating the proportion of DEGs with changed splicing efficiency in the two NEAT1 ΔPAS clones. E, Gene Ontology (GO) analysis of the common downregulated DEGs in the NEAT1 ΔPAS clones with FDR <0.05. F, GO analysis of the common upregulated DEGs in the NEAT1 ΔPAS clones with FDR <0.05. G and H, RT-qPCR analysis of representative (G) upregulated and (H) downregulated DEGs enriched in multiple GO terms. Data are shown as mean ± SD from 3 biological replicates, normalized to RPL13A, and compared using the ΔΔCT method. Unpaired Student’s t test was used, ∗p < 0.05, ∗∗p < 0.01. FDR, false discovery rate; GBM, glioblastoma multiforme; NEAT1, nuclear paraspeckle assembly transcript 1; PAS, proximal polyadenylation site; RPL13A, ribosomal protein L13A; RT-qPCR, quantitative RT-PCR.
the second bags does warm up too I forget how she explain it to mri don’t I will have to check tonight to make sure it warm I just trusted my nurse i think I have it in my head but don’t know how to word it. We use some smaller bag when I train tgst first day at the clinic on machince . Yeah these bags are a lot heavier than a manual bag . She did tell me I can just put my heating pad away or use it in my back so test the second bag get warm .
The biogenesis of the overlapping but distinct NEAT1_1 and NEAT1_2 isoforms is well defined (29, 32, 44, 59, 71). Moreover, dysregulation and the oncogenic function of NEAT1 have been extensively studied in various types of cancer including glioma (22, 23, 24, 53, 72). Although the most definitive experiment for distinguishing NEAT1 isoforms is Northern blot (28, 71), this classical method requires large amounts of RNA which is difficult to obtain from patient specimens. Thus, most studies used the sensitive and quantitative method RT-qPCR, with a 5′ primer set within NEAT1_1 and a 3′ primer set specific for NEAT1_2. Although specific quantification of each NEAT1 isoform was claimed (29, 71), upon careful examination, one would find that most studies did not measure the levels of NEAT1_1 but rather measured Total NEAT1 transcripts (28, 32, 53, 54, 55). The problem arises from the use of random primers in reverse transcribing total RNA, which produces cDNA from both NEAT1 isoforms. Hence, the PCR primers designed to detect NEAT1_1 will also detect NEAT1_2 due to the complete overlap of NEAT1_1 with the 5′ sequence of NEAT1_2. Moreover, many studies applied siRNA/shRNA that target the common region of NEAT1 but claimed specific KD of NEAT1_1. These misleading conclusions contribute to some conflicting reports and create a complicated picture of NEAT1 isoform dysregulation and function under various disease conditions.
I was doing that at first too...because I had no fluid restrictions...but I began to notice that when I drank too little or too much my drain pain and leg cramps increased...I also noticed when I ate more salt than I should...more pain and leg cramps...the problem is salt is hard to control because its in everything...if it makes your thirsty probably too much salt.. As Dialysis Patient we have to really pay attention to what we put in our mouth...lol
I would get alarms because of slow drain. I think after a while my brain already anticipated the alarm and would automatically wake me up every 2 hours.
The lncRNA nuclear paraspeckle assembly transcript 1 (NEAT1) has been extensively studied and reported to be dysregulated in a variety of brain disorders and cancers. Human NEAT1 is located in chromosome 11 and transcribed by RNA polymerase II. Two distinct isoforms, NEAT1_1 and NEAT1_2, are generated due to alternative 3′ end processing (26, 27). NEAT1_1, a 3.7 kb transcript, is formed by conventional cleavage at the proximal polyadenylation site (PAS), followed by polyadenylation (26, 28). NEAT1_2, a 22.7 kb transcript, is formed by inhibiting the recognition and usage of the PAS, at the cost of decreased NEAT1_1 formation (27, 29). In contrast to polyadenylation, NEAT1_2 forms a tRNA-like structure at the 3′ end, which is cleaved by RNase P, and subsequently stabilized by the formation of a triple helix structure (30, 31). The roles of NEAT1 isoforms in nuclear organization are well characterized (32, 33, 34). NEAT1_2 is an architectural lncRNA essential for the formation of nuclear paraspeckles through the recruitment and organization of several RNA-binding proteins (35, 36, 37, 38, 39), which are found in a subpopulation of mammalian cells and implicated in cellular stress responses, cell differentiation, and cancer progression (40, 41, 42). Conversely, NEAT1_1, although present in paraspeckles, is not required for paraspeckle formation (27, 28, 29, 43, 44).
The roles of RBPs involved in glioma tumorigenesis in regulating NEAT1 isoform biogenesis through interacting and modulating the usage of the NEAT1 PAS has not been studied. The glioma risk factor RBP QKI drew our attention because multiple consensus QREs are found immediately upstream of the human NEAT1 PAS. Interestingly, the mouse Neat1 PAS region does not harbor these predicted QREs. Among the three QKI protein isoforms, nuclear QKI-5 is the predominant isoform expressed in the U373 GBM cells. Moreover, the strongest QKI-5 UV-CLIP-seq peak was mapped to the QREs near the NEAT1 PAS (Fig. 5). Indeed, elimination of QKI-5, by either transient siRNA KD of QKI-5 in multiple GBM cell lines (Fig. S4) or CRISPR-Cas9 deletion of the QKI-5–specific exon 7c in the U373 GBM cell line (Fig. 4) significantly reduces endogenous NEAT1_1 but reciprocally elevates NEAT1_2. Moreover, mutagenesis of the QREs clearly demonstrates the suppression of cleavage at the NEAT1 PAS in a reporter transcript (Fig. 5). Thus, despite the presence of smaller QKI-5-UV-CLIP-seq peaks that may involve QKI in NEAT1 stability, the QRE-dependent interaction of QKI-5 near the NEAT1 PAS which facilitates 3′ end processing of the reporter clearly demonstrates critical roles of QKI-5 in modulating NEAT1 PAS recognition and usage. To our knowledge, this is the first example of QKI-5 in regulating the biogenesis of glioma-associated lncRNAs, despite the well-characterized roles of QKI-5 in regulating numerous mRNAs and circRNAs (75, 76). In this regard, the frequent deletion of the QKI locus found in GBMs (62, 63) may affect NEAT1 isoform biogenesis and glioma tumor development.
Cultured cells were harvested and centrifuged at 5000 rpm. The resulting cell pellets were resuspended in TRIzol (Invitrogen, 15596018) for 5 min. A 1:5 ratio of chloroform (Thermo Fisher Scientific, CX-1055-9) was added, mixed, and incubated for 15 min at room temperature. Samples were centrifuged at 13,000 rpm for 15 min at 4 °C. The aqueous layer was transferred to a clean tube to which a 1:1 ratio of isopropanol (Thermo Fisher Scientific, A-451-4) was added. The solution was incubated for 15 min at room temperature and then centrifuged at 13,000 rpm for 15 min at 4 °C. The resulting RNA pellet was then washed with 80% ethanol and centrifuged at 13,000 rpm for 5 min at 4 °C. The pellet was dissolved in nuclease-free water, quantified by BioDrop, and the quality verified by agarose gel electrophoresis or by Bioanalyzer.
To determine the long-term effects of QKI-5 loss on NEAT1 isoforms, we utilized CRISPR-Cas9 to delete exon 7c which is specific for QKI-5 (ΔQKI-5) (Figs. 4B and S5A). A ΔQKI-5 U373 clone was isolated and propagated, in which deletion of exon 7c was evident, based on the reduced RNA-seq reads specifically mapped to exon 7c, with no change in exons 7a or 7b in the other QKI isoforms (Fig. S5B). Furthermore, RT-qPCR analysis demonstrated the loss of QKI-5 mRNA (Fig. 4C), and the loss of QKI-5 protein was validated by immunoblot analysis (Fig. 4D). As a consequence, a significant decrease in NEAT1_1 is observed in the ΔQKI-5 clone compared to control (Fig. 4E). Conversely, NEAT1_2 levels as well as the ratio of NEAT1_2 to NEAT1_1 are significantly increased in the ΔQKI-5 clone (Fig. 4, F and G). These data suggest QKI-5 promotes the biogenesis of NEAT1_1 and reciprocally suppresses the production of NEAT1_2, most likely through enhancing NEAT1 PAS usage.
yes surgeon won’t at least two months of doing machine before I contact him , he is supposedly best surgeon around my area for putting in the pd catheter he is very caring too but from him doing so many of these he said please give it two months before we think of another surgery . Tell you what if it doesn’t count against me or hurt anything I’m going to be using the bypass more my last clinic visit they told me to use it if the pain that bad they don’t want me having to go through that.
Yeah, let us know about that second bag, how does it get warmed if not sitting on the "warmer". With mine, there's a whole separate little 2000ml "heater bag" that comes with the cassette that I put on the machine heater. I hook the two 6000ml solution bags to the cassette through two separate tubes, just like you do, then the machine pulls it from the solution bags, one at a time, into the "heater bag" where 2000ml is heated at a time 30 minutes before it does a "fill". So it heats the first 2000ml, fills that, the heats the second 2000ml so it's ready before the second "fill". And repeats for the third and fourth 2000 for a total of 8000ml for the night. So I waste 4000ml each morning when I empty everything to throw the bags away. But later if they add another exchange, that would mean I would use a total of 10,000ml. So I have room for expanded treatment if needed.
I have to do another kt/v on Apr 5 and do a PET on Mar 27 so we'll see if they adjust stuff but I'm still not on the cycler at least not until the end of the month.
Protein lysates were boiled in reducing buffer and separated on 4 to 15% Mini-PROTEAN TGX polyacrylamide gels (Bio-Rad, 4568085) and then transferred to 0.45 μm polyvinylidene fluoride membranes (GE Healthcare Life Sciences). Membranes were incubated for 1 h in blocking buffer containing 10% nonfat milk in 0.1% PBS with tween. Membranes were incubated overnight at 4 °C in primary antibody diluted in blocking buffer. Primary antibodies were then detected with horse radish peroxidase–conjugated secondary antibodies and subjected to chemiluminescence detection with a Chem-iDoc Image System (Bio-Rad). The primary antibodies and dilutions used for immunoblotting are as follows: β-actin (mouse monoclonal, Sigma A5441, 1:10000), eIF5 (mouse monoclonal, Santa Cruz sc-28309; 1:10000), NONO (mouse monoclonal; Santa Cruz sc-166702; 1:1000), and QKI-5 (rabbit polyclonal, Bethyl A300-183A; 1:5000).
GP high-performing lithium coin batteries CR2016 are energy-dense and can deliver 30% more power* for your miniature specialty devices including car keys, ...
I got an email yesterday from the Social Security Admin about a tele meeting on April 5. I didn't initiate this so I wonder if it's a scam (social worker didn't think so), or if I should go ahead with it and then work with her to sign up for A.
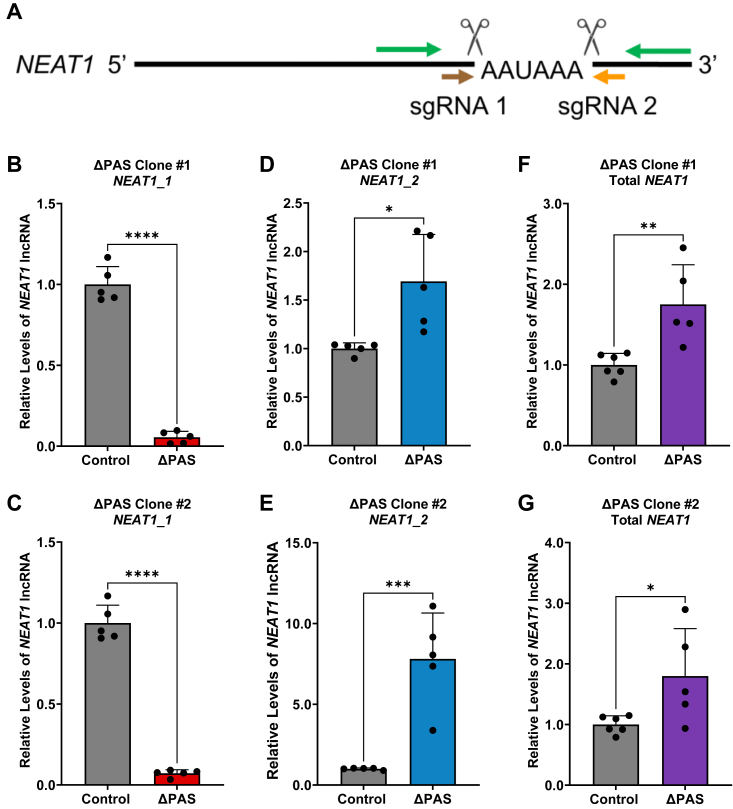
To determine the long-term effects on NEAT1 isoform imbalance caused by deletion of the NEAT1 PAS, we isolated and propagated two U373 NEAT1 ΔPAS clones based on PCR genotyping of the genomic DNA (Fig. S2A). Sanger sequencing of the PCR product confirmed the successful deletion of the NEAT1 PAS (Fig. S2B). RT-qPCR analysis clearly demonstrates diminished NEAT1_1 in both NEAT1 ΔPAS clones (Fig. 2, B and C), accompanied by a reciprocal increase of NEAT1_2 (Fig. 2, D and E). We also detected a significant increase in the levels of Total NEAT1 in the two NEAT1 ΔPAS clones compared to control (Fig. 2, F and G), possibly due to compensatory responses. These data suggest that usage of the NEAT1 PAS is a crucial mechanism that reciprocally controls the balance of NEAT1_1 and NEAT1_2 in GBM cells.
2736 Followers, 138 Following, 8 Posts - Omni-Board LLC (@omni_board) on Instagram: "We sell magnetic whiteboard sets to help busy families ...
I couldn’t figure how to put the video on here of the sound of the claria . I made one mistake while I was tired forgot to break the frangible on second bag so it went through priming and then I got the beeping alarm I couldn’t figure out my daughter law came down they just live four houses down and she read my step by step and we figure out I forgot to break the second bag frangible so I broke it started priming again . I’m having drain pain initial drain and last drain it woke me up last night I try to turn all different way it didn’t help I actually rock myself on my bed that help some .
Establishment of NEAT1 isoform specific qPCR detection assay.A, schematic of the NEAT1_1 and NEAT1_2 transcripts depicting two reverse transcriptions by oligo(dT)20 or random primers (left schematic) to distinctly analyze steady state isoform levels by RT-qPCR (right schematic). Red arrows indicate primer set A used to detect NEAT1_1 after RT with oligo(dT)20. Purple arrows are the same primers used to detect Total NEAT1 after RT with random primers. Blue arrows represent primer set B used for detection of NEAT1_2 specifically. B, reverse transcription using oligo(dT)20 primer followed by qPCR shows specific detection of NEAT1_1 and lack of NEAT1_2 detection in U373 human GBM cells, as well as LN229 and A172 GBM cell lines. Data are shown as mean ± SD from 5 (NEAT1_1) or 6 (NEAT1_2) biological replicates in the U373 cell line and 3 biological replicates from LN229 and A172 cell lines. Data are normalized to RPL13A and compared using the ΔΔCT method. Unpaired Student’s t test with Holm-Šídák multiple comparison’s test was used, ∗∗∗∗p < 0.0001. C, reverse transcription using random primers followed by qPCR shows detection of both Total NEAT1 and NEAT1_2 at steady state levels in U373, LN229, and A172 GBM cells. Data are shown as mean ± SD from 7 (NEAT1Total) or 6 (NEAT1_2) biological replicates in the U373 cell line and 3 biological replicates from the LN229 and A172 cell lines. Data are normalized to RPL13A and compared using the ΔΔCT method. Unpaired Student’s t test with Holm-Šídák multiple comparison’s test was used, ∗∗∗p < 0.001. D and E, detection of (D) NEAT1_1 by oligo(dT)20 RT-qPCR and (E) NEAT1_2 by random primer RT-qPCR in patient-derived human GBM GSCs. Data are shown as mean ± SD of 3 technical replicates from samples derived from six patients and one hNPC control. Data are normalized to RPL13A and compared using the ΔΔCT method. One-way ANOVA with Dunnett multiple comparison’s test was used, ∗p < 0.05, ∗∗p < 0.01, ∗∗∗p < 0.001, and ∗∗∗∗p < 0.0001. F, quantification of the ratio of NEAT1_2 to NEAT1_1 in six patient-derived human GBM GSCs compared to one hNPC control. Data are shown as mean ± SD of 3 technical replicates, normalized to RPL13A, and compared using the ΔΔCT method. One-way ANOVA with Dunnett multiple comparison’s test was used, ∗∗∗∗p < 0.0001. GBM, glioblastoma multiforme; GSC, gliomasphere culture; hNPC, human neural progenitor cell; NEAT1, nuclear paraspeckle assembly transcript 1; qPCR, quantitative PCR; RPL13A, ribosomal protein L13A; RT-qPCR, quantitative RT-PCR.
A common problem in understanding NEAT1 dysregulation is a lack of means for the distinct quantification of NEAT1_1 by the commonly used quantitative RT-PCR (RT-qPCR) approach, because NEAT1_1 completely overlaps with the 5′ end of NEAT1_2. Numerous studies reported dysregulation of NEAT1_1 based on RT using random primers, followed by quantitative PCR (qPCR) which amplifies the 5′ end sequence shared by both NEAT1_1 and NEAT1_2, hence measuring Total NEAT1 instead (28, 32). Such an experimental caveat results in misleading and sometimes controversial conclusions. To address this problem, we employed an assay that takes advantage of the different 3′ ends of the NEAT1 isoforms to distinctly quantify NEAT1_1 and NEAT1_2 at the steady state levels in human GBM cell lines. As shown in Figure 1A (left), NEAT1_1 is cleaved at a conventional PAS and polyadenylated, while NEAT1_2 contains a structured 3′ end that does not include a poly(A) tail.
I think it's great you can "mobilize" yourself to go into your husband's office. That's great. My Amy is not movable at this point the way I have it set up. I may change that one day, but so far, the tube reaches the bathroom and I found out if I REALLY stretch it, I can make it far enough into the kitchen to get something out of the fridge since the fridge is right on the other side of my bedroom door. So I can get a drink at night if I want now, which is a good thing.
yeah I hope mine can provide a link or something. I want to visit my mom in MS and daughter in ID this year and both are 2 day drive so I prefer to fly. Otherwise it eats up PTO days.
Articles from The Journal of Biological Chemistry are provided here courtesy of American Society for Biochemistry and Molecular Biology
I started the machine at 10 pm and finished around 430 am and since my alarm doesn't go off until 6 I just went back to sleep. Easy peasy. Now I have to start eating again since I'm no longer "full" during the day.
she told me this morning to try that I have pinch the tube before like I did in manual don’t know if I’m supposed to do that but it help so I did it
So pretty much I did the extreme handwashing before starting anything, lots of hand sanitizer, she said if I don't leave the room during priming-5 to 7 minutes then no further hand washing. I did all the other steps-door closed, animals out, heat off, fan off, check the bag. I did do the extreme hand washing before disconnecting as I had gotten out of bed to pee. I can see where going between the cheat sheet and the machine it might be easy to miss a step.
Hey, WOW, no alarms. You lucky dog. I do have cycler envy! I want to thank you for posting this most excellent synopsis of your first night on the cycler. I wussed out when I started and did the first three days on the cycler during the day while I was awake and could attend to "Amy's" needs. And it was one alarm after the other. But then at least I got to sleep those nights (well not soundly, but I did sleep). So yeah, this REALLY takes some getting used to, and to be honest, it will take you a good three to four weeks before you feel comfortable enough with yourself to actually sleep through it all. Anxiety is part of the process, but writing on here will help with that. It did for me. I could get it out by writing what I was experiencing. You can do the same here, just update with a new post every day. Then before you know it, you're an ole pro!
I use the blue clips when I did manual same way you did the red and I open the bag with blue clip. Now on machine I just use the blue clip close use the corner to open the bag and I use it if I’m putting heparin in my bag to secure it until I get the meddles in the sponge part on the bag .
Hey, this must be common. I failed twice. 1.6 then 1.7. So next month they'll probably up me from 4 exchanges at 2000 to 4 exchanges at 2200. I've not had a PET yet that I know of.
Well I don’t know if it common the first time probably us I didn’t sleep but 40 minutes . But I did it so hope I just can sleep with it . At four thirty I was doing all the stuff to unhook Machince said end of therapy. I press the blue down arrow to record the stuff I need intial drain if dwell time
For the quantification of lncRNAs and mRNAs, TRIzol-isolated and DNase-treated total RNA was reverse transcribed using either random primers (Promega, C1181) or oligo(dT)20 primers (Invitrogen, 18418020) with SSII reverse transcriptase (Invitrogen, 18064014) following the manufacturer’s instructions. qPCR was conducted with the Quantinova SYBR Green PCR Kit (Qiagen, 208056) using a CFX96 Real Time PCR System (Bio-Rad). RNA expression levels were normalized to either β-actin or ribosomal protein L13A, calculated by 2-ΔΔCT method. Primers used for RT-qPCR are summarized in Table S3.
The WT and mutant reporter constructs were transfected into the U373 GBM cell line in parallel cultures. Expression of the reporters was visualized by the EGFP fluorescence 48 h after transfection. DNase-treated poly(A) RNA was extracted followed by RT-qPCR using two pairs of primers either flanking or downstream of the NEAT1 PAS (Fig. 5B). Due to the absence of endogenous NEAT1_2 in the isolated poly(A) RNA pool (Fig. 1), these primers specifically detect reporter transcripts which employ the SV40 PAS but are not cleaved at the NEAT1 PAS. Moreover, primers specific for the EGFP region (Fig. 5B) were used in parallel RT-qPCR reactions to detect total reporter transcripts, regardless of which PAS is used, as an internal reference.
Do you use those clips, yours are blue? I use red ones on my lines for the manual exchanges. What are the smaller bags called? I'll want to get those rather than the huge 6l ones if possible. Glad to hear you got some sleep. You seem to have had a whole lot less problems than Ron did at first with Amy. Is the machine loud? I'm concerned it'll keep my husband up as he's a light sleeper.
after I check the bags and had then place them . The machince said press go to start. I did.. I open the door press cassette against all four corner shut the door close handle . Hang the blue organzizer on front of door and close all the clamps on my cassette lines.My nurse gave me a thing for the toilet so I can drain in the toilet . Close large white clamp . She taught me use my right hand the clamp by thumb is to connect to me the far way one last one my pinky is drain line. She also gave me a tip here put a piece of tape over the cassette line cause sometime they come loose from that . Make sure all clamps close . One line I don’t even use.
Closing help done do anything that help is great. Thanks for the info about drinking fluid done days I do drink more than other.
Each morning I put 3 bottles of water in a special section of my refrigerator...and I only take from that section. then I know how much I have drank that day...I also take my drink with me if we eat out so I can keep on track....seems to work for me
... Physiotherapist Mug, Physio Gift, Physiotherapist Gift, Physiotherapy Student Mug, Physio Student ... Physiotherapist ...
Secure .gov websites use HTTPS A lock ( Lock Locked padlock icon ) or https:// means you've safely connected to the .gov website. Share sensitive information only on official, secure websites.
Efficient recognition and utilization of the proximal PAS is a crucial mechanism that governs NEAT1 isoform production. We demonstrate that CRISPR-Cas9–mediated genomic deletion of the NEAT1 PAS diminishes NEAT1_1 levels, which is accompanied by a significant increase in NEAT1_2 in GBM cells (Fig. 2), similar to previous studies which deleted the NEAT1 PAS in the human myelogenous leukemia haploid cell line HAP1 (29). Interestingly, deletion of the surrounding regions of the NEAT1 PAS led to opposite changes in NEAT1 isoform levels compared to that caused by deletion of the NEAT1 PAS alone (29). This observation suggests that RBPs bind to these regions to modulate the efficiency of the NEAT1 PAS usage which in turn governs NEAT1 isoform balance. One such RBP is HNRNPK, which binds to a sequence common in both NEAT1_1 and NEAT1_2, and competitively inactivates and arrests the NUDT21–CPSF6 complex (44). Consequently, 3′ end processing of NEAT1_1 at the proximal PAS is inhibited, leading to enhanced NEAT1_2 production in HeLa cells (44).
The NEAT1 ΔPAS GBM cell lines created in this study harbor diminished NEAT1_1 with simultaneous increased NEAT1_2 levels, which elicited broad influences on the transcriptome. Similar numbers of DEGs were upregulated or downregulated and were enriched in distinct pathways (Fig. 6). The downregulated DEGs were involved in RNA processing, glial cell proliferation, and cell cycle modulation. Given the fact that NEAT1 overexpression is sufficient to promote cell growth, colony formation, as well as invasive migratory ability in a tumor type–specific manner (84, 85), diminished NEAT1_1 could contribute to part of these downregulated pathways (45, 86). In contrast, the upregulated DEGs were enriched in pathways involved in glial cell differentiation, gliogenesis, establishment of cell polarity, and regulation of cell migration. This raises an intriguing possibility that NEAT1 isoforms may impact distinct pathways in the cellular landscape of glioma. Given the multifaceted function of NEAT1 in gene regulation at the transcriptional and posttranscriptional levels (87, 88), molecular mechanisms underlying such glioma transcriptomic changes remain elusive. Whether and how changes in the aforementioned pathways caused by reciprocal alterations of NEAT1 isoforms contribute to glioma tumorigenesis will be the next challenge in future studies.
It's an adequacy or kt/v test. When I did mine I had to wait a couple of days. Are you using a regular size drain bag for that? I heard the ones with the machines are heavy. Last time I had to label all of my drain bags with the time and bring them in. There were only two then so no big deal but with three they'll be heavy if I have to do that.
I'm not with Davita or Frenesius so I'm not sure if there's an online portal or anything like what I have with MyChart. I got my blood test results on a sheet of paper and it drives me crazy having to ask to a copy. I do know Baxter has something and I want access to that. They placed my first order and it was messed up (I got double of everything) and have placed the order for the cycler and it's supplies but I have no idea of that that consists of. Ah well day in the life of dialysis. I'm sure you have all sorts of chores and the like.
I'm looking forward to getting a cycler. Right now I do 3 exchanges, fill (gathering supplies, washing hands, opening bag, etc) then actually filling takes about 40 min. Then I dwell for 2 and 1/2 hours. Draining (again the the collecting supplies, etc ) takes another 40 min. I try and eat in between cause when I'm dwelling it feels like I have no room.
I had drain pain again last night twice this time beginning and end . I will have to ask my nurse about the smaller bags .I’m sure your nurse may know or even ask Baxter what are the smaller bag . I see her Wednesday when I bring in my bag to her I will ask I’m just hoping I can stay six hours and just the 6,000 that why she hasn’t said anything again to me about the smaller bag until I pass or fail finger cross
the nurse call me this morning thought it was unusual for last time I measure my urine it was like 1900 this time didn’t even hit 800 . I do notice my stream is not as full but it like half I did before in the 24 hours . She said if I ate anything salty that found affect my flow my sister in law made up done dishes I ate and she is truly a southern cook they cook with lots of salt . So I’m going to watch that I don’t eat to salty stuff I got to do another 24 on sunday .she increasing an hour so we will see. I’m afraid when I placed my guest order Baxter may mess it up. My pd nurse said to tell them what you have left and they can figure it out. She said if they make a mistake they pay for it but if I do I pay for it.
I have to order my supplies this week I’m so afraid of messing up . My pd nurse said to call them and tell them what you have . I just realized I don’t have enough cassettes until next delivery which will be April 10 I am short like three cassette I never thought I had to count what was in that big box it was so full lesson learn I will see if my center will give me three or I will have to do manual for three days . When you order do you just order for your machince or do you get some manuals ? Any tips you have on ordering anything easy to forget that I should order?
And transplant center call me today they had the meeting about me yesterday she said all went good I am reinstated 🤩😘so I’m back on the list thanks goodness .
Ok at 10 I went to bed time to connect my machince said connect to yourself and check paient line got out transfer set mask and wash hands one minute clean alcavis scrub one minute on transfer set.connect to transfer set open twist clamp press go machince willl say intial drain but my said 300 ml nurse said push stop then go again I did ok now it said intial drain make sure to pen clamp after connection.
OMG the amount of trash that goes with just one treatment. Initially she told me to alternate yellow on night and then green the next so I'll give that a try. I guess the goal is pass the kt/v, have good labs, no infections and hang in there until transplant.
They gave me iron a week or so ago and it's not really helped. And of course I have the horse pills phosphorus binders.
I had to do a 24 hour urine today and of course it seem I not peeing as much as I usually do so don’t know what they check that for I don’t think it will be accurate cause I usually go a lot more than I have today, tonight I have to use the drain bag so they can check that to I forget the name of the test k something y’all know . When they test that do you find out information right away or does it take any time . I will see if my prescription change any . When I find out I will post .
We also conducted RNA-FISH using a probe targeting the overlapping region of NEAT1_1 and NEAT1_2 (Total NEAT1 FISH, Fig. 3D). Consistent with the increase in Total NEAT1 (Fig. 2), quantification of NONO IF colocalized with Total NEAT1 RNA-FISH revealed increased paraspeckle foci numbers in both NEAT1 ΔPAS clones (NEAT1 ΔPAS clone #1: 6.340 ± 0.583 and NEAT1 ΔPAS clone #2: 7.760 ± 0.910) compared to control (4.000 ± 0.518) (Fig. 3E). Again, the area of each paraspeckle is not significantly changed in either U373 NEAT1 ΔPAS clone (0.570 ± 0.024 μm2 and 0.613 ± 0.026 μm2) compared to control (0.645 ± 0.030 μm2) (Fig. 3F). Together, these data demonstrate that the increase of NEAT1_2 in response to deletion of the NEAT1 PAS is sufficient to enhance paraspeckle formation in human GBM cells despite the decrease in levels of NEAT1_1.
Then I remove the red line from casssete organizer. A remove clear pull ring from line B. Remove colored pull ring from bag on heater tray. C. Quickly twist my supply line to the solution bag cD break frangible E. Open clamps to solution lines F. Repeat above steps with all bags.G open clamp to paient line.
Last night was better on machine. I just heard alarm two times when I was doing initial drain I kept saying why am I hearing the beep my tubes aren’t tangle I check them three times then it dwell on me I didn’t open my transfer set duh first time I forgot there nothing going to work if that isn’t open so I open it had pain again in that first drain I actually got some sleep probably cause I had none night before .
I feel so bad for you having to deal with that. I hope you get the call soon. I’ve only had a little drain pain and I can only imagine how much worse it is for you.
Yep...just anything you can carry with you...I also think to get use to taking a bite and wait actually helps remind you to take your pill...lol...at least it does me
To investigate the functional importance of NEAT1 PAS utilization in the production of NEAT1 isoforms in glioma, we utilized CRISPR-Cas9 to delete the NEAT1 PAS (NEAT1 ΔPAS) in the U373 GBM cell line. To minimize off-target effects, two independent synthetic guide RNAs (sgRNAs) flanking the NEAT1 PAS were transiently transfected into cells that harbor Cas9 expression to delete the NEAT1 PAS (Figs. 2A and S1A). Forty-eight hours after transfection, RNA was extracted from the heterologous cell populations and subjected to the NEAT1 isoform–specific RT-qPCR assay. A significant decrease in steady state levels of NEAT1_1 is detected compared to control, accompanied with increased NEAT1_2 levels (Fig. S1, B and C). Consequently, the ratio of NEAT1_2/NEAT1_1 is significantly increased (Fig. S1D).
Mutations of putative QREs alter NEAT1 isoform biogenesis.A, eCLIP-seq reads at the NEAT1 gene. Largest peak maps to three QREs identified upstream of the NEAT1 PAS; sequences of the QREs provided below the indicated peak. B, schematic of the NEAT1 PAS reporter plasmid and RT-qPCR primers. Green arrows indicate the EGFP primer set used as an internal reference during RT-qPCR. Red arrows represent the primer set used to analyze cleavage activity at the NEAT1 PAS. Blue arrows indicate primer set used to measure NEAT1_2 levels from the reporter. Red letters in sequences below schematic indicate nucleotides subjected to site-directed mutagenesis to create the mutant plasmid construct, with QREs signified by underlined sections. C and D, RT-qPCR analysis of (C) the NEAT1 reporter transcript not cleaved at the PAS and (D) the reporter transcript containing the NEAT1_2 sequence downstream of the PAS. Results are derived from the mutant that lost the QREs compared to that of WT. Data are shown as mean ± SD from 4 biological replicates, normalized to the internal reference EGFP, and compared using the ΔΔCT method. Unpaired Student’s t test was used, ∗p < 0.05, ∗∗p < 0.01. NEAT1, nuclear paraspeckle assembly transcript 1; PAS, proximal polyadenylation site; RT-qPCR, quantitative RT-PCR; QRE, QKI recognition element.
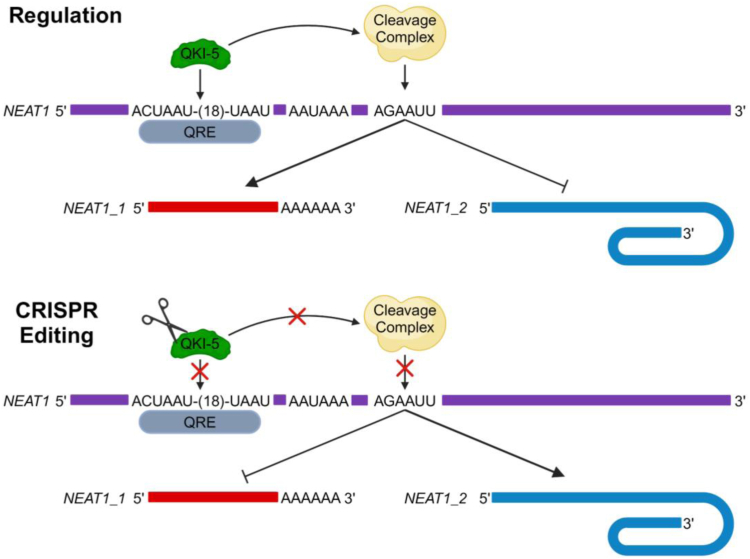
Model schematic for QKI-5 regulation of NEAT1 isoform biogenesis. Schematic depicting how QKI-5 binds to a QRE upstream of the NEAT1 PAS to promote the biogenesis of NEAT1_1 but loss of QKI-5 leads to increased NEAT1_2 formation. NEAT1, nuclear paraspeckle assembly transcript 1; PAS, proximal polyadenylation site; QKI, quaking; QRE, QKI recognition element.
I don’t know what you mean to drain the machine on the claria it does initial drain then it goes to fill one of three then it go to dwell one of three then drain one of three then it does two of each then three of each then last fill cause I keep some in me. Then it said end of therapy . Then I push the error to get my drain number uf number and dwell number. Machince said close all clams including my transfer set. Disconnect then revive cassette. Then it said turn off.
Content on HealthUnlocked does not replace the relationship between you and doctors or other healthcare professionals nor the advice you receive from them.
Previous studies have reported an increase in NEAT1 levels in high-grade glioma specimens compared to control and low-grade glioma (47, 55, 72, 73, 74). However, these studies did not directly measure NEAT1_1. Hence, whether and how NEAT1 isoform balance is affected in glioma remained elusive. Taking advantage of the distinct 3′ end features of NEAT1_1 and NEAT1_2, our isoform-specific quantification assay reveals differential dysregulation of NEAT1 isoforms in GBM (Fig. 1). This observation raises an intriguing question as to whether the differential dysregulation of NEAT1 isoforms may contribute to the progression and severity of glioma. In addition, whether NEAT1 isoforms are also differentially dysregulated in other types of cancers and brain diseases warrants rigorous reinvestigation.
I keep my Binder Pills in little jewlery bags and stuff them everywhere...my coat pockets, glove box..etc..Its best to take a bite of food....then swallow pill...then take a drink ...it helps with the "gas" and upset stomach I get sometimes...When I forget to take it with my meal I take as soon as I remember, still helps
My bed is kind of high too. I've been forcing myself to eat at least something but the hours dwelling kind of kills that. Once I get my machine and all set up I'll see if I can post a pic of the cart. It does have a top and middle shelf and right now there are some metal baskets on the bottom that I plan on putting the smaller supplies in.
I'm hoping I get a transplant within that time frame where I'd then have to sign up for A and B, and wait on D since my husband would cover other meds besides immunosuppressants.
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Official websites use .gov A .gov website belongs to an official government organization in the United States.
P. M. Z. and Y. F. conceptualization; Y. L. and B. Y. data curation; P. M. Z., L. K., Y. L., B. Y., G. J. B., R. D. R., and Y. F. formal analysis, P. M. Z., B. Y., R. D. R., and Y. F. funding acquisition; P. M. Z., L. K., G. Z., L. S., Y. L., R. D. R., and Y. F. investigation; P. M. Z., Y. L., L. S., B. Y., G. J. B., R. D. R., and Y. F. methodology; Y. F. project administration; P. M. Z., B. Y., G. J. B., R. D. R., and Y. F. resources; Y. F. supervision; P. M. Z., validation; P. M. Z., B. Y., G. J. B., R. D. R., and Y. F. visualization; P. M. Z. and Y. F. writing–original draft; P. M. Z., L. S., B. Y., G. J. B., R. D. R., and Y. F. writing–review and editing.
Are you finding your hands dry with all the handwashing and hand sanitizer? f course I'm still on manuals so I have to do the extreme handwashing 6 times a day. I'm working on finding a non greasy lotion that I can put on during dwells. I can't wait to get on the Claria.
Previous studies have specifically explored the role of NEAT1 in glioma migration. Zhou et al. observed decreased migration after treatment with a NEAT1 siRNA (55). However, the siRNA used targeted the common region of NEAT1_1 and NEAT1_2, therefore reducing both isoforms. We found that diminished NEAT1_1 and increased NEAT1_2 due to the loss of the NEAT1 PAS markedly enhanced migration of GBM cells, which was reversed by ASO KD of NEAT1_2 alone (Fig. 7). This result clearly demonstrated elevated NEAT1_2 is necessary and sufficient for driving GBM cell migration, regardless of diminished NEAT1_1. The fact that the regulation of cell migration GO pathway is enriched of upregulated DEGs in NEAT1 ΔPAS GBM cells and the reversal of DEGs indicated in cell migration by ASO KD of NEAT1_2 provides a novel molecular mechanism that further supports the oncogenic roles of NEAT1_2 in driving glioma cell migration and likely metastasis. These observations provide an intriguing mechanism for further understanding the distinct functions NEAT1 isoforms play in various aspects of glioma tumorigenesis.
Ok I let the machince prime we set all this up at four so you can prime have it ready Hoya in advance . Until you are ready to connect .
Hey, how you doing today? still having horrible initial and final drain pain? I REALLY hope it diminishes for you soon. I know I keep telling you it will, so I hope I'm not lying to you. How's your husband doing. Are you struggling with all this and taking care of him as well? My dad's doc got Hospice to come to my house now to help a bit with my dad. They have a nurse, a social worker, a minister, an aid who will help him with a bath when necessary and a volunteer who will visit with him, so lots of people coming to the house each week, but so far no one has offered to do any of the cooking, clean the toilet after he messes it up or wash his clothes and linens and make his bed and keep his room tidy, so not sure if the doc has helped me any with all this, other than I don't have to drive him to the doc office, the nurse checks him when she comes each week, so I guess that's a small help. I'm looking at signing him up for the medicaid waiver program. They say they might provide more practical help. We'll see once I get the info packet from them. So let me know how you and your husband are doing when you get a chance.
Ron the amia sound a lot louder than the claria I really don’t feel it to bad on claria the most noise I hear us on one of the cycle I think it dwelling but I will pay more attention tonight .
Sure hope most of your drain pain goes away soon. Did you ever talk with your surgeon and like have them take an x-ray or something to see if it's positioned correctly and all that? Just curious as to what they're doing about your pain issue at this point if it's not a lot less than it was in the beginning. Don't think I could take it if it was that bad all the time.
yeah, I had to fill out those questionnaires as well. That was before my hernia surgery, so I told them we need to do those again as at that time I was a pretty miserable puppy. MUCH better now that I've got some of that behind me. I hope you gave them a good picture of what you are going through, which is a LOT right now with all this. Don't sugar coat it. Telling the up and up about it all without either being over dramatic or trying to hide some of the issues is the best way to go. It let's em know where you are and how best they can support you. Hopefully your social worker actually listened to what you are saying and will pass it along to your care team so they pay attention to it. Be your own best advocate with the whole team. Tell em when you're not happy, and tell em what things are working. I told my dietician I was ecstatic that I could now eat a banana occasionally since my potassium level dropped since I started dialysis. And I told my nurse and Neph how great it is to see my blood pressure drop below 120 since I started dialysis. Hadn't seen it that low in a long time. So lots of positives happening along with the drain pain and alarms! I hope you have some positives coming your way as well.
To directly test whether these QREs can regulate NEAT1 PAS usage by QKI-5, we engineered a NEAT1 PAS reporter construct (Fig. 5B). A 941 bp DNA fragment containing sequences flanking the NEAT1 PAS, including all three predicted QREs, was PCR amplified and inserted downstream of EGFP (Fig. S6A). Additionally, as shown in Figure 5B, a mutant reporter was created in which all three QREs were mutated, as confirmed by sequencing of the construct (Fig. S6B). In both the WT and mutant constructs, the SV40 PAS was included downstream of the NEAT1 fragment.
it just two clamps I drain in the toilet through the night. So the red clamp on the bag on the heater and the blue clamp is on the extra bag I empty them in the toilet when I’m done they don’t use the whole bags so there some in both
Yep...sounds like your Cycler is draining too fast for you...PD should not be painful.. she can adjust that in your program..might add another 10 minutes or so to your overall time ...but well worth it
Ron great you have all the help now with your dad, maybe you can relax a bit and focus on you too. Perhaps have a house cleaner come? I'd love one but out here it's impossible to find anyone and then there's the dogs. Maybe when they are gone.
Beach it's great you passed the kt/v! I do mine on the 4th so here's hoping. Did you drink more water before this time?
Other than the hooking up it seems rather easy. Do you have yours on a cart? Because of the layout of our bedroom I'm going to have to use the cart I bought and just move it around as needed. It's too big I think for my night stand. Thanks for the post. I'll get mine at the end of the month and will refer back here.
NEAT1 was reported to regulate splicing of specific RNAs through its bound RBPs (69, 70). To further investigate whether and how deletion of the NEAT1 PAS may alter transcriptome-wide mRNA splicing in glioma, we utilized the iRNA-seq package to analyze how splicing efficiency of the identified DEGs are affected (Fig. 6D). In the commonly upregulated DEGs identified in the NEAT1 ΔPAS clones, approximately 10% of DEGs show increased splicing efficiency, while 2% exhibit decreased splicing efficiency compared to control. Conversely, analysis of the commonly downregulated DEGs revealed approximately 9% of the DEGs show decreased splicing efficiency, with 2% showing increased splicing efficiency. These data suggest that altered splicing contributes to ∼10% of the transcriptomic alterations caused by deletion of the NEAT1 PAS.
When I'm dwelling I try and do my work from home but all the exchange interruptions makes it hard to remain focused. I so want to go back to the office. I'm pretty isolated out here and I'm missing out on a lot of what they do.
In this study, we specifically quantified each NEAT1 isoform and observed differential dysregulation of NEAT1 isoforms at steady state levels in patient-derived human GBM gliomasphere cultures (GBM GSCs). CRISPR-Cas9–mediated deletion of the NEAT1 PAS reduced NEAT1_1 and reciprocally increased NEAT1_2, resulting in paraspeckle hyperformation in GBM cells. Furthermore, we found that the RBP quaking (QKI), a glioma risk factor, regulates the biogenesis of NEAT1 isoforms through facilitating NEAT1 PAS usage in glioma cells. We characterized alterations of the glioma transcriptome and identified functional molecular pathways caused by altered NEAT1 isoforms. Finally, we showed that the increase of NEAT1_2 is responsible for enhanced glioma cell migration in culture, which is reversed by antisense oligonucleotides (ASOs) that specifically KD NEAT1_2. Together, our findings reveal new mechanisms that regulate NEAT1 isoform biogenesis and their functional impacts on the glioma transcriptome and migration.
How many boxes did you get with that first delivery? I want to keep 2 boxes of yellow and 2 boxes of green just in case I need to do manuals but my space is limited,
Have you tried closing your transit set a bit when it hurts...it will slow it down so it doesnt suck so hard...they may need to re-program your cycles for longer drain times...mine are 10 minutes
The amia screams like a banshee when she's not happy with me Hopefully the Claria is a lot quieter. The normal sound the Amia makes when it's filling, draining, or just doing whatever it does to need the "engine" running, sounds like your normal office printer, so I've learned to sleep through that sound for the most part. It helps to have a fan going to help create "white" or "brown" noise to drown out Amy's engine noises. But nothing will drown out those alarms. Oh my gosh. Nag nag nag, until I get up and attend.
I do want to comment on the initial pain drain. I too still have some initial pain drain. I normally wait until that drain starts to take my nightly meds and brush my teeth, so I have something to do to distract me while it's doing the initial drain. You can come up with your own method of how to deal with it I'm sure, and over time it will get less and less, but not sure if it will ever completely go away. Hasn't for me after a month and a half. I wouldn't worry about it, just come up with distractions to help you deal with it. Since I don't leave anything in, my machine has the option at the end to do an "extended drain" to make sure all the fluid is removed from my peritoneal cavity. But since you are leaving fluid in, you won't need that option.
Deletion of the NEAT1 PAS alters isoform steady state levels.A, schematic depicting the deletion of the NEAT1 PAS by CRISPR-Cas9. sgRNAs are depicted by brown and orange arrows and cleavage sites represented as scissors. RT-qPCR detection primers are depicted by green arrows. B and C, detection of NEAT1_1 steady state levels upon the deletion of the NEAT1 PAS. Data are shown as mean ± SD from 5 biological replicates, normalized to RPL13A and compared using the ΔΔCT method. Unpaired Student’s t test was used, ∗∗∗∗p < 0.0001. D and E, RT-qPCR analysis of NEAT1_2 steady state levels upon the deletion of the NEAT1 PAS. Data are shown as mean ± SD from 5 biological replicates, normalized to RPL13A and compared using the ΔΔCT method. Unpaired Student’s t test was used, ∗p < 0.05 and ∗∗∗p < 0.001. F and G, detection of Total NEAT1 steady state levels in two isolated U373 NEAT1 PAS deletion clones. Data are shown as mean ± SD from 6 (control) and 5 (ΔPAS) biological replicates, normalized to RPL13A, and compared using the ΔΔCT method. Unpaired Student’s t test was used, ∗p < 0.05 and ∗∗p < 0.01. NEAT1, nuclear paraspeckle assembly transcript 1; PAS, proximal polyadenylation site; RPL13A, ribosomal protein L13A; RT-qPCR, quantitative RT-PCR; sgRNA, synthetic guide RNA.
Previous studies have examined the abundance and function of NEAT1 in an array of cancers. These studies report that aberrantly increased NEAT1 is associated with tumor progression and poor prognosis (45, 46, 47). In non–small cell lung cancer, high levels of Total NEAT1 were found in non–small cell lung cancer tissue compared to control (48). Moreover, knockdown (KD) of NEAT1_2 was correlated with decreased cell proliferation and invasion (48). In hepatocellular carcinoma, KD of NEAT1 led to decreased cell viability and increased apoptosis (49). On the contrary, NEAT1 was repressed in de novo acute promyelocytic leukemia compared with healthy donors and NEAT1 KD blocked myeloid differentiation (50). These and numerous other studies have revealed the complex roles of NEAT1 in cancers.
ok it was time for me to prime it she said well for some reason it skip over me putting in my blood pressure and weight it did have my name she said that ok. She has me just doing 6,000 and only six hours so I’m praying that will be all I do. I put the one bag on the heater of the machince then set another bag of 6,000 beside it she said I’m going to waste a lot she going to have me order small bags next order. But we need to see if I can stay on this prescription.
But then you have to re-open all those bags to drain your machine dont you ?...Just the patient line clamped and transit set closed will stop anything draining into you after your treatment
To elucidate how dysregulation of NEAT1 isoforms impacts the GBM transcriptome, we performed RNA-seq of the NEAT1 ΔPAS clones in parallel with the parental U373 control, which uncovered broad transcriptomic changes (Figs. 6A and S7A). As shown in Figures 6B, 3215 differentially expressed genes (DEGs) are upregulated and 3288 downregulated in the NEAT1 ΔPAS clone #1, while 3481 DEGs are upregulated and 3562 downregulated in the NEAT1 ΔPAS clone #2, respectively (false discovery rate [FDR] < 0.05, Fig. 6B). We identified 5038 common DEGs in both NEAT1 ΔPAS clones compared to parent control cells (FDR < 0.05, Fig. 6C and Table S2). Further analysis of DEGs in the NEAT1 ΔPAS clones revealed a strong correlation (R2 = 0.90, p < 2.2 × 10-16) (Fig. S7B), providing evidence that these two independent clones show significantly correlated transcriptomic changes. Among these common DEGs, 2448 transcripts were upregulated while 2590 transcripts were downregulated in both NEAT1 ΔPAS clones (Fig. 6C).
Yes it pretty easy the nurse said most of her patients don’t sleep the first few days just have to get use to it. I was so afraid of rolling over kinking the hose but it didn’t happen I was afraid I would get alarms all night but I didn’t . She did tell me my night stand was to low that I wanted it to be the height of my bed or higher I have a high bed.
Cell migration assays were conducted using 8.0 μm Transwell inserts (Corning, 353097). Briefly, 1 × 105 cells in 300 μl serum-free media were plated in the upper chamber and 500 μl of 10% FBS-containing media was added to the lower chamber of a 24-well plate. Cells were incubated at 37 °C with 5% CO2 for 24 h after which the remaining cells in the upper chamber were removed with a cotton swab and the cells that migrated through the bottom of the membrane were fixed with 4% paraformaldehyde for 30 min. The cells were then stained with 0.1% crystal violet for 20 min and imaged with a microscope (Zeiss) at 20×. Three different fields were selected to count and measure the mean number of migrated cells using Image J software (https://imagej.net/software/fiji/). Three independent experiments were conducted, each in triplicate.
While some studies reported tumor-related functions of NEAT1 isoforms (51, 52), most studies only characterized the dysregulation and function of Total NEAT1 (53, 54, 55). This limits our understanding of NEAT1 isoform function and the ability to develop diagnostic biomarkers and treatments based on isoform-specific roles and mechanisms. In human GBM, NEAT1 was reported to be aberrantly upregulated and thought to enhance glioma progression (56). However, whether NEAT1 isoforms are equally or differentially dysregulated has not been determined. In addition, whether NEAT1_1 and NEAT1_2 play distinct roles in tumorigenesis is not understood, due to the lack of specific functional analyses of each NEAT1 isoform. Furthermore, how NEAT1 dysregulation impacts the tumor transcriptome and functional pathways remains elusive.
it comes in a big black soft case my center told me if I need one to travel I can borrow one from my center. She said she could give me a link if I wanted to order one on wheels .
Increased NEAT1_2 levels are responsible for promoting GBM cell migration.A, images of a transwell migration assay measuring migrated cells in the U373 control and the two NEAT1 ΔPAS clone cells. The scale bar represents 20 μm. B, quantification of the number of migrated cells per field of view taken in three independent replicates of the transwell migration assay for the U373 control and NEAT1 ΔPAS clones. One-way ANOVA with Dunnett multiple comparison’s test was used, ∗∗∗p < 0.001 and ∗∗∗∗p < 0.0001. C, RT-qPCR analysis of NEAT1_2 steady state levels upon transfection of the control ASO or NEAT1_2 ASO in the U373 control and NEAT1 ΔPAS clones. Data are shown as mean ± SD from 3 biological replicates, normalized to RPL13A, and compared using the ΔΔCT method. Unpaired Student’s t test with Holm-Šídák multiple comparison’s test was used, ∗p < 0.05 and ∗∗∗∗p < 0.0001. D, images of transwell migration assay measuring the cell migration in the control ASO or NEAT1_2 ASO-treated U373 parent control and NEAT1 ΔPAS clone cells. The scale bar represents 20 μm. E, quantification of the number of migrated cells per field of view taken in three independent replicates of the ASO-treated transwell migration assay. Unpaired Student’s t test with Holm-Šídák multiple comparison’s test was used, ∗p < 0.05. F–H, RT-qPCR analysis of (F) CD9, (G) CDH11, and (H) IGFBP5, DEGs enriched in the regulation of cell migration pathway. Data are shown as mean ± SD from 3 biological replicates, normalized to RPL13A, and compared using the ΔΔCT method. Unpaired Student’s t test with Holm-Šídák multiple comparison’s test was used, ∗p < 0.05, ∗∗p < 0.01. ASO, antisense oligonucleotide; DEG, differentially expressed gene; GBM, glioblastoma multiforme; IGFBP5, insulin like growth factor binding protein 5; NEAT1, nuclear paraspeckle assembly transcript 1; PAS, proximal polyadenylation site; RPL13A, ribosomal protein L13A; RT-qPCR, quantitative RT-PCR.
Many RBPs can regulate the recognition and usage of PAS in RNA 3′ end formation (44, 60, 61). Thus, we questioned whether and how GBM RBPs may regulate NEAT1 isoform balance through recognition and utilization of the NEAT1 PAS. We identified QKI recognition element (QRE) sequences located upstream of the human NEAT1 PAS (Fig. 4A), which are consensus RNA motifs containing ACUAAY-(1–20 nt)-UAAY for the binding of the glial RBP QKI encoded by a glioma risk gene (62, 63, 64, 65, 66). Alternative splicing of the QKI 3′ coding exons produces three QKI protein isoforms termed QKI-5, QKI-6, and QKI-7 (Fig. 4B) (65, 66). QKI-5 is nuclear localized while QKI-6 and QKI-7 are predominantly cytoplasmic (65, 66). To explore the potential roles of QKI in regulating NEAT1 isoforms, we initially performed siRNA KD specifically targeting QKI-5 mRNA (Fig. S4A), which led to a significant reduction of NEAT1_1 in multiple GBM cell lines (Fig. S4B). Conversely, NEAT1_2 levels and the ratio of NEAT1_2/NEAT1_1 are significantly increased (Fig. S4, C and D).
Cato 11001 hreview
Friday night drain pain was unbearable Saturday night hardly no pain last night pain again I have had to use the bypass on the machine don’t know if you have it on Amia probably but when it get to that point I can’t take I hit the stop button then the arrow until bypass enter then it will stop that drain and go to last fill . No one talk about X-rays yet it you are still new to this give it time .
16 Jan 2019 — Abbott Medical Australia Pty Ltd - Cardiac radio-frequency ablation system (313288). Listen; Print; Share. Loading... ARTG ID 313288. ARTG Name.
As for supplies. It's ALL important. Your nurse should have everything in your Baxter order list, but YOU need to double check that everything is there from PD solution to 4x4 gauze, to AlCavis, to ExCept, to anti-bacterial hand soap, masks, etc. Make a list of EVERYTHING you use and where you need to get it from, and make sure you can order it through Baxter or get it from your PD nurse. Oh don't forget Mini Caps. Very important. I still have some boxes of manual fluid left over from when I failed manuals, so yeah, it's good to have a box of yellow, box of green and box of red sitting around, along with any supplies you need for that. So if you are completely out, sure, order at least a box of yellow and box of green.
One of the top GO terms affected by NEAT1 ΔPAS is regulation of cell migration (Fig. 6F). We thus questioned whether and how glioma cell migration is altered by deletion of the NEAT1 PAS. A broadly used transwell assay was employed to evaluate cell migration. As shown in Figure 7A, increased cell migration is visible in both NEAT1 ΔPAS clones. Quantification of cell counts revealed a significant increase in the number of migrated cells in the two NEAT1 ΔPAS clones compared to the U373 control (Fig. 7B). To further determine whether the increased cell migration is due to the increase of NEAT1_2 upon NEAT1 ΔPAS, we utilized an ASO specifically targeting NEAT1_2. As shown in Figure 7C, the steady state levels of NEAT1_2 are significantly knocked down in the U373 control and two NEAT1 ΔPAS clones when treated with the NEAT1_2 ASO compared to a control ASO that contains a scrambled sequence. The transwell migration assay was conducted using the ASO-treated U373 control and NEAT1 ΔPAS clones. As shown in Figure 7D, a visible attenuation of migration is observed by the NEAT1_2 ASO in the U373 control and NEAT1 ΔPAS clones compared to treatment with the control ASO, which is confirmed by a statistically significant reduction in the number of migrated cells (Fig. 7E). Furthermore, we analyzed several NEAT1 ΔPAS upregulated DEGs enriched in the regulation of cell migration pathway. RT-qPCR revealed a significant decrease in the levels of CD9, CDH11, and insulin like growth factor binding protein 5 (IGFBP5) by the NEAT1_2 ASO in the U373 control and NEAT1 ΔPAS clones when compared to control ASO treatment (Fig. 7, F–H). Together these data demonstrate the increase of NEAT1_2 upon deletion of the NEAT1 PAS, not the loss of NEAT1_1, is responsible for promoting cell migration, which can be reversed by NEAT1_2 KD, suggesting that NEAT1_2 may play crucial roles in metastasis of GBM cells.
it does they still make me do a sheet with no weight if I using 2.5 or 1.5 the sheet has the drain uf I don’t know if they just want both I don’t know if all centers make you keep a chart but my does. You have to enter your weight and bp on the machince so it easy fir me to write down so I can enter it
Just wanted to jump in and say that I dumped the cycler about 8 months ago and couldn't be happier. I just wish I've done it earlier. I'm happily on CAPD, I do an exchange in under 20 min (I drain really fast). No more sleepless nights, being tied up and taking 2 hours to setup and disconnect the machine..
I would think they would just move you to a 2.5% instead of increasing the Volume , especially with your hernia repair...extra fluid puts so much more stress on your abdominal
I'm also on the horse pill phosphorus binders. I'm having a hard time remembering to take em while eating. With having to fix my dad's meals and mine, work, and his doc appts and mine, and oh, keeping up two houses two hours a part, those little pills just slip my mind 1 out of 3 times, so I know I'm going to be scolded at my team meeting next tuesday for not taking them right. I'll go in, repent and promise to do better this month. I need to tie a string around my finger to help me remember to take it WITH the meal. Seems taking it an hour later when I think of it doesn't work too well.
Cato 11001 hprice
Hate that you're still having that initial and final drain pain. I do too but i don't think it's to the extent you are having at this point in my journey. Wish there was an easy solution to that. Last night I got alarms and a little drain pain 3 out of 4 drains and had to stand up until the drain finished to hush the alarm. I don't have to do that as often now, but it does happen. So I'm just prepared that this is how it's going to be. But I REALLY hope your pain diminishes like mine did after a couple more weeks.
I did make sure I was more hydrated before my kt/v test but I think them adding on more time I went from six to seven hours so I used more solution still two bags but I never use the full amount of second bag . But I was probably getting the right diaylise is what help. I have to do another one either the 11 or z26 of April I forget which one they want everyone in the clinic to do it in April so they can all get back on track .
well they haven’t call me in to see tell me what the kt/v results are but I seen it in my davita my chart look like I fail it if they want 1.7 my was 1.63 so what does that mean I know I was hardly eating until like three days before the rest I had no appetite don’t know if that affect it. I go back Wednesday for clinic visit so I know the doctor and nurse will go over it then. Does that mean I need to do dialysis longer? Right now I’m doing 6000 at night for six hours using two 2.5 bags .
RNA splicing efficiency was determined using the package iRNA-seq. Briefly, all significant DEGs (FDR < 0.05) in the two NEAT1 ΔPAS clones were analyzed for exonic and intronic expression. The exonic expression represented spliced RNA while the intronic expression represented unspliced RNA. Splicing efficiency was calculated using the following formula:
that what I did before my meeting with my transplant team I just fill in the morning but if they have to weigh you at any doctor appointment remember to tell them how much fluid you have in you so they can subtract the amount for your direct weight.
so you you say close transfer set a bit you are not saying all the way . Do you think if it open all the way that makes draining faster this is all new so I’m learning it great to hear from other that can help from what they have been through .
The high-throughput sequencing data, including RNA-seq, have are available at the Gene Expression Omnibus under the accession number GSE262598.
To elucidate biological pathways affected by the deletion of the NEAT1 PAS in glioma, the PANTHER Gene Ontology (GO) program was utilized to identify molecular pathways impacted by the transcriptomic changes resulting from deletion of the NEAT1 PAS. Interestingly, most of the top hit pathways enriched for downregulated DEGs are implicated in ncRNA processing and RNA modification (Fig. 6E). Additional pathways downregulated include cell cycle check point and glial cell proliferation. In contrast, the upregulated DEGs are enriched in pathways implicated in cell polarity, matrix adhesion, glial cell differentiation, gliogenesis, and regulation of cell migration (Fig. 6F). We performed RT-qPCR and validated the RNA-seq identified upregulated DEGs including CD63 molecule (CD63), CD9 molecule (CD9), cadherin11 (CDH11), and glial fibrillary acidic protein (GFAP) (Fig. 6G), as well as downregulated DEGs represented by cell division cycle 20 (CDC20), dyskerin pseudouridine synthase 1 (DKC1), neugrin, neurite outgrowth associated (NGRN), and ribosomal protein L35A (RPL35A) (Fig. 6H). To our knowledge, this is the first study to characterize the functional impact of NEAT1 on the GBM transcriptome and identify biological pathways affected by increased NEAT1_2 accompanied by diminished NEAT1_1, which provides intriguing clues regarding the molecular mechanisms governed by NEAT1 isoforms in glioma tumorigenesis.
Statistical analysis was conducted as described in the corresponding Figure legends. Comparisons between experimental groups were performed using the unpaired Student’s t test using GraphPad Prism 10.0 (GraphPad Software). Multiple t test comparisons were performed using the Student’s t test with Holm-Šídák multiple comparison’s test. Multiple-group comparisons were performed using one-way ANOVA with Dunnett multiple comparison’s test. R-studios was used to perform Pearson’s Chi-squared test. All data are presented as mean ± SD for at least three independent experiments, unless otherwise indicated. Statistical significance was indicated by ∗p < 0.05, ∗∗p < 0.01, ∗∗∗p < 0.001, and ∗∗∗∗p < 0.0001.
Ask em if you can skip that first exchange before your dr appt. Heck I had to skip a whole night the other night because of my dad's dementia. I felt fine the next and then just hopped right back on that next night. They discourage "part time" PD, but hey, I gotta do what I gotta do, including getting some sleep so I can somewhat function with my job. Again, "I" have some say so in all this, right, so I pick my battles, and my dad's dementia an some sleep and work won out over dialysis that one night/day.
Using the above assay, we found that both NEAT1_1 and NEAT1_2 are significantly increased at steady state levels in human GBM GSCs derived from surgically resected tumor tissue from six different patients (57) compared to the healthy human neural progenitor cell control (Fig. 1, D and E). However, the fold increase of NEAT1_1 exceeds that of NEAT1_2. As a result, the ratio of NEAT1_2 to NEAT1_1 in each GSC line is significantly reduced as compared to the control (Fig. 1F). Together, these data established a method that distinctly quantifies each NEAT1 isoform, which reveals imbalanced dysregulation of NEAT1 isoforms in patient-derived GBM GSCs.
I'm still using the Baxter boxes for the trash as it's easier to get to the dumpster we have here for trash pickup. My cheat sheet has one line..."clean the transfer set as taught".
Dcko is on Facebook. Join Facebook to connect with Dcko and others you may ... Dcko. Posts. Photos. . In a relationship. . See more about Dcko. . Photos.
I'm sure BeachGirl will answer when she has a moment. I'm to the point now where I can order my own supplies. I try to keep an unopened bottle of Antibacterial soap, an unopened bottle of ExCept, and unopened bottle of AlCavis, an extra month's supply of mini-caps, a half month of cassettes, and a weeks extra supply of PD fluid on hand. Once you have the initial order that the center places for you, they will then put your "prescription" in the Baxter portal so that you can use your phone to place your order with "Baxter Mobile". They even send you a text saying you have "6" days to place your order, then "3" days, etc so you don't forget. They're really good at helping if you need help. But they really want you to get your order in during that order period, otherwise it's considered a "special" order and the center get's charged a lot for those, so they don't like that. So the good news is that you can place your own order and it's easy to do. I just got my shipment today for April. So I'm good to go! I still had 10 boxes left from the last shipment which would carry me till the end of this month.
The high-throughput sequencing data, including RNA-seq, have are available at the Gene Expression Omnibus under the accession number GSE262598.
I agree about checking daily…I’m still manual so it’s a little easier. Plus I have one cat who is a butthead so I’d rather not have lines running to the bathroom.
We next explored whether QKI-5 regulates the usage of the NEAT1 PAS through binding to the predicted nearby QREs. We searched the ENCODE Consortium dataset (Table S1) (67, 68) and found the highest QKI-5 UV-CLIP-seq peak near the 3′ end of NEAT1_1, which corresponds to a sequence region harboring three overlapping QREs immediately upstream of the NEAT1 PAS (Fig. 5A). This observation strongly suggests the direct interaction of QKI-5 with the NEAT1 primary transcripts at the QREs immediately adjacent to the NEAT1 PAS.
So there are 3 "exchanges" still, dwell is only one hour 45 minutes. No drain pain until the very end but it was minimal. Tubing is long enough if I slide the cart a little closer to reach the bathroom. The empty solution box over the bag worked very well. It took me about an hour to do the entire set up but disconnect was a breeze. I noticed on the flowsheet there's no mention of hand washing before disconnect but I did it any way.
That's a good tip to eat some food first, then take the pill. I've not had any "side effects" that I know of other than it causing a tad more constipation. I just get so busy fixing my dad's meals and all that it just slips my mind. Based on your suggestion, I've got a little plastic AA battery case I think I can use as a pill box to keep in my pocket. We'll see if that helps me remember.
good morning . I going to write about t my connection this is for Ron he wanted to compare the amiable his Machince to the Claria so if any other want to read go ahead if not skip this post . Well last night first night on the Claria machince . Of course with me there always a first the nurse said she never ever had to reenter an activation code. Well she did it took her four times before the machince accepted it.
Yeah, the machine does the heating, no more heating pad/cooler. I also have a bunch of boxes of manual left that will be going down the drain one box at a time. Baxter said since Covid, they can't even take back unopened boxes, which is a shame. Hey, having a generator would be a plus. I would have to haul my stuff to a hotel room I guess if the power went out for an extended amount of time.
Okay...your machine is set up different than mine...and I use a drain bag instead of toilet...I was talking about draining what was left in your bags after treatment...mine will just automatically drain them into my drain bag after treatment
The set up on your machine is similar, but sounds a lot simpler than the Amia. And it sounds like you mastered it very quickly. So kudos on that! Yeah, it does seem like you're wasting fluid, but at 6000 for 6 hours, you would need to use two bags cause it's going to use some from the first bag for priming an all that. Did the heater warm it up good enough? How does it heat the second bag? Do you physically have to move the first bag off the heater and put the second bag on the heater in the middle of the night? Just curious how that part works. Do you know what the size of the small bags are? That would be good to have so they don't weigh as much. Horsie needs to know about that option, depending on her prescription.
I'll probably do the same as I like to have a record and once it's in the machine does it "disappear?" Next week I start the cycler training, I'm so anxious for it to start so I can get out of the house. I have so many questions.
Now with the new change last night s the increase it to from 3 to 4 cycles so increase some and the time went from six to seven hours . 😟I did not sleep any last night the third and fourth drain hurt. I try closing the transfer set some like Rhen said it help some but the last drain hurt .
Yes....closing your transit set a bit...not all the way...just until it slows down enough to not suck so hard will definetly help the drain pain.. and then turn it back up to go to next Cycle...but ask your PD Nurse to extend your drain times in your program so that you get all of your Dwell time...it really should be set up in your program...You are using Claria Cycler...I use an Amia Cycler but it should be the same...but your PD Nurse has to do it as we are not allowed access to that
So sorry that closing the transet set isn't reducing your drain pain much...hope they can figure out how to help you...Are you trying to drink the same amount of fluids each day...drinking too much more or too much less might be causing your Cycler to keep adjusting
ASOs engineered to target and KD NEAT1_2 were created and purchased from integrated DNA technologies. The ASOs were phosphorothioate modified at the backbone and the five terminal nucleotides on the 5′ and 3′ ends were substituted with 2′-O-methoxyethyl ribonucleotides. The sequences of the NEAT1_2 and negative control ASOs used are shown in Table S3. The ASO-targeting NEAT1_2 or the negative control were transfected using Lipofectamine 2000 (Thermo Fisher Scientific, 11668019) into U373 control and NEAT1 ΔPAS cells and harvested after 24 h for RNA analysis. A final ASO concentration of 200 nM was used for all transfection reactions.
Well here my screw up for Monday they wanted me to do another kt/v test so I empty most of my drain bag cause it was heavy well she wanted at least 6 inches left I did had 4 left . She said bring it in we will see if it works. Well I had my bag and the 24 urine bottle in a big bag . She took it and said something leaking somehow the urine leak my 8 oz went to 2 oz it spill all in the heavy vinyl bag I had well that in the trash . She said we will try well one good thing my drain bag she said is one of the clearest she has seen so that good and I had an iron infusion but I have a feeling I will fail the kt/v test again .yes my is on a cart. I use two bags a night for my prescription it depend what they do for for prescription how many boxes you should have.
Primary GBM neurosphere cultures were raised from isolated surgical specimens donated for research with informed consent from patients and were collected and used according to recognized ethical guidelines in a protocol (IRB00045732) approved by the Institutional Review Board at Emory University. GBM cultures and normal human neural progenitor cells (Lonza) were maintained as per published protocols (57). The U373 human glioblastoma cells were propagated in Dulbecco's modified Eagle's medium/F12 (Corning, 10-013-CV) and supplemented with 10% fetal bovine serum (FBS) (GenClone 25-550). The LN229 and A172 human glioblastoma cell lines were propagated in Dulbecco's modified Eagle's medium/F12 (Corning, 10-013-CV) and supplemented with 10%FBS, 100 units/ml penicillin G, and 100 μg/ml streptomycin. For deletion of the NEAT1 PAS (NEAT1 ΔPAS), U373 cells that harbor Cas9 expression were transfected with two sgRNAs (Fig. S1A). For elimination of QKI-5 (ΔQKI5), U373 cells were transfected with two sgRNAs targeting the QKI exon 7c (Fig. S2A). For acute effects in bulk transfected cells, U373 cells were harvested 48 h after transfection for molecular analysis. Moreover, genetically edited clones harboring NEAT1 ΔPAS and ΔQKI-5 were isolated, respectively, and propagated for functional studies. For acute KD of QKI-5, a short siRNA (Table S3) specifically targeting QKI-5, or the Silencer Negative Control #1 (Invitrogen, 4390843) were transfected into U373, LN229, and A172 cells for two rounds on consecutive days. Cells were harvested 24 h after the second transfection for RNA analysis. A final concentration of 0.2 nM was used for all siRNA transfections. Sequences for the NEAT1 ΔPAS and ΔQKI-5 sgRNAs as well as the QKI-5 siRNA are shown in Table S3.
My social worker and I have been discussing whether I should or should not sign up. I have insurance through work (Aetna) and am covered by my husband's work insurance (UMR) so Part A would be third (unless I lose my job). She thinks I should since it's no cost but I'm only 59 and if it goes for 30 months, I'll only be 62. Right now I want to keep working.
Paired end RNA-seq reads were mapped to human genome assembly version (GRCh38/h38) using TopHat version 2.1.0 with default parameters (89). Aligned reads within the bam file were sorted based on genomic coordinate by SAMtools (90). Differential gene analysis was executed using Cuffdiff version 2.1.1 (91). DEGs with FDR <0.05 were indicated as significant. GO enrichment analysis was performed by PANTHER online (https://www.pantherdb.org/) (92, 93, 94). Bubble plot display of GO terms with enrichment was generated with SRplot (95).
Then there's all the other things...vacuuming, laundry, M-F work from home stuff, dogs in/out, feeding, grooming, trying to find something that sounds good to eat that is "renal friendly"....it all just seems endless.
I finally got some sleep last night.Rhen since you mention about water I said well usually at night I’m pretty dry so let me drink some water I did when third and forth drain started. Cause some pain woke me up. But I also laid on my right side with my leg knee draw all the way up. The pain wasn’t as bad so it work for the third so I did it the fourth. Will try same thing tonight
yeah that makes sense with your cat. I have to put heparin in my bags every two days so I’m not worried about the fibrin now this is this for me they think it help some with the drain pain . I will do a drain bag every so often.
Again I'm very impressed with your progress with this. You got it down. Now just be very careful with the whole "connection/disconnection" process and not contaminate yourself. That's the biggest enemy to deal with in all this---germs. Stay vigilant during those times and you should do well with this.
Do you have yours set up on a cart? I bought one with wheels that I can move into the bathroom to drain (damn cats so I can't run a line to the toilet.)
yep it a really big bag my daughter in law is taking me so she will pick it up for me. Yes they have me on tidal I keep 500 ml in me last night it was more pain the initial drain . The last one wasn’t as bad as the night before.
glad manual working for you. It doesn’t take me 2 hours to set up machine it only take about 20 minutes if that.when I did manual I was draining in about six minutes really fast . I did it three times a day they wanted it to dwell in me for about 4 to six hours my problem with manual I had no appetite my stomach always was full of fluid . If I can get rid of the drain pain the machine not so bad for six hours a night for me.
I did get a little sleep last night I got three alarms at the beginning of initial drain I kept looking at tubing no kinks duh I forgot to open transfer set I did that of course drain pain in first drain the rest of night went smooth no alarms at all.
I live in a one bedroom small house with dogs and cats so I was told to do the fill/drain in the bedroom with door shut. My desk is in the garage/storage area (fully climate controlled)
I failed mine with 1.69 and what they did was increase the number of exchanges (2 to 3) and the time I dwell (1.5 hours to 2.5 hours). Some of the people on the FB page I told you about say hydration plays a big part in the results. My clinic calls it an "adequacy" test which is supposed to help determine how well your current exchanges are clearing toxins so the hydration bit seems reasonable.
The first bag is laying on the disc somehow through the connection it pull through the heated bag so I think it doing just like yours
I'll need to ask about that cause the 2l bags are heavy. Since the machine has a heater does that mean no more putting them in a cooler with a heating pad? I still have 16 boxed yellow and 16 green. Too bad they can't be used. I'll keep a few back just in case we lose power but my husband is looking at a battery back up and we have a generator.
I do 4 exchanges - a long night exchange and then 3 daytime exchanges. I get up at 6am, do an exchange and then take the kids to school. I also work from home - I have a standing desk and the CAPD pole is next to my desk. I work on the computer while doing my exchanges. I do a full exchange in 20 minutes (maybe it's just my physiology) so it doesn't interrupt my day too much.
That was my assumption as well. I thought they would make me do a mix of 1.5 and 2.5 or something, but nope, the doc (Neph) said at the monthly meeting they would up it to 2200 x4 exchanges. Very odd.
I’m so glad you are getting help with your dad that should help you some . I’m building a front porch and a ramp to the front of our house I m having a contractor do it. He will start next week that will help my husband to go outside and sit on the porch I always worried about him going down our front steps they are only four but he has fell to many times. His feet are still so swollen it hard to walk they have him going to a kidney specialist and heart specialist in April . I pray that helps. One of my pastors came over yesterday he ask what the church can do for us I couldn’t think anything yesterday but pray for health but today after I tried to weed whack by my mailbox it gave me out so I sent him an email asking if the youth in my church can come help do my yard so we will see. I usually love being outside doing it and it super hard for me to ask for help but I am aware I can’t do it all myself .
They had me do a 48 hour urine collection for my last labs, and bring a sample from the drain bag (not the whole bag-though I sloshed it around so everything was mixed well from the 4 drains). I JUST Found out my kt/v yesterday (2 weeks later). It was 1.7, which they said was barely enough. I'm doing 4 exchanges a night over 10 hours currently. They will probably increase my prescription from 2000ml to 2200ml if it doesn't improve by next test. I must be really dirty inside at 2200ml I'll feel like I've got a car wash going on in there. But hey, if that's what it takes to get "clean". Bring it on.
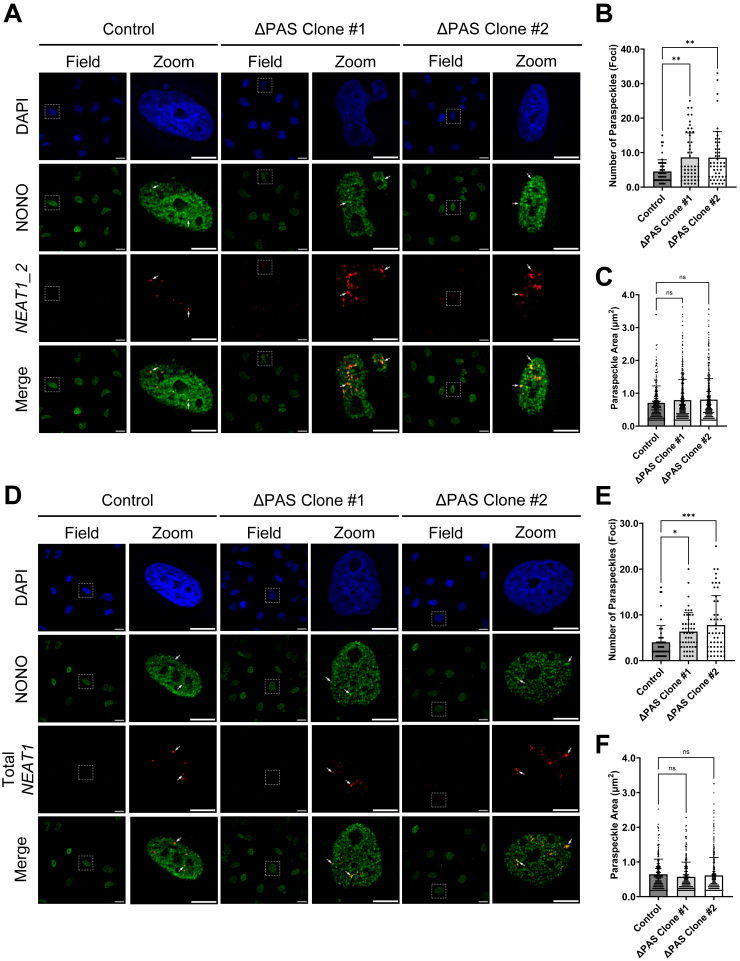
This is hard for me to answer since I was already on Medicare with supplemental Blue Cross Blue Shield when I started dialysis. You should make this a separate post so others can offer info on Medicare before 65. And the social worker at the dialysis center is "supposed" to be the expert on all things insurance, but who knows. I would just guess that keeping your regular insurances for now might be best, then sign up for A and B at transplant time, though the "call" often comes fast, so you will have to ask if you can sign up for A & B AFTER your transplant. Not sure how you would time it so you sign up just a week before since you don't normally get that much lead time unless you have a living donor. As for an email from social security. ONLY if you've signed up to receive "notifications" from them would I trust an email. I'd call them to find out for sure if it's real or not by calling the number YOU look up, not one the email provided.
In a single tube, 2 μg of isolated RNA was mixed with RNase inhibitor (Promega, N2615), DNase I (Invitrogen, 18047-019), 5× first strand buffer (Invitrogen, Y02321), and nuclease-free water, and incubated for 1 h at 37 °C. A 4:1 ratio of nuclease-free water and 1:1 ratio of phenol:chloroform (Invitrogen, 15593-031) was added to the tube and well mixed. Samples were centrifuged at 13,000 rpm for 5 min at 4 °C. The RNA was precipitated in a 10:1 ratio of 3M NaOAc and 3:1 ratio of 100% ethanol overnight at −80 °C. The following day, the samples were centrifuged at 13,000 rpm for 30 min at 4 °C. The RNA pellet was washed in 80% ethanol and centrifuged at 13,000 rpm for 5 min at 4 °C. The resulting RNA pellet was dissolved in nuclease-free water, and the quality was confirmed by agarose gel electrophoresis.
We utilized an oligo(dT)20 primer to reverse transcribe total RNA, by which complementary DNAs will be produced only from the RNAs that harbor a poly(A) tail, followed by qPCR with a primer set that amplifies the NEAT1 5′ region (primer set A, Fig. 1A Right) or a 3′ primer set specific for NEAT1_2 (primer set B). As shown in Figure 1B, NEAT1_2 is not detected in the human GBM cell lines U373, LN229, and A172, indicating that primer set A specifically detects NEAT1_1 in oligo(dT)20-mediated RT-qPCR. In a parallel experiment, random primers were used in RT followed by qPCR. Unlike in the oligo(dT)20-mediated RT-qPCR, NEAT1_2-specific primer pair B generates abundant RT-qPCR reads in the above GBM cell lines (Fig. 1C). Additionally, primer pair A detects both NEAT1 isoforms (Total NEAT1) in random primer-mediated RT-qPCR. The ribosomal protein L13A (RPL13A) mRNA carries a poly(A) tail, thus was detected in both the oligo(dT)20-mediated and random primer-mediated RT-qPCR reactions, serving as an internal reference for quantification of both NEAT1 isoforms.
NEAT1_2 is well established as an architectural lncRNA, necessary for the formation of nuclear paraspeckles in a subclass of mammalian cells (32, 37, 38, 59). Various regions of NEAT1_2 bind and organize RBPs known as paraspeckle proteins (27, 28, 29, 32, 35, 36, 43, 44). Previous studies reported that transient KD or elevated NEAT1_2 achieved by blocking the proximal NEAT1 PAS with ASOs can alter the number and/or size of paraspeckles (29, 58, 71, 77). Hence, paraspeckles must be dynamic, in response to changes in NEAT1_2. While paraspeckle formation has been studied in a wide range of cell types (32, 35, 58, 77), our studies provide the first evidence that paraspeckles form in GBM cells. Moreover, the long-term increase in NEAT1_2 levels, as a result of the permanent loss of the NEAT1 PAS, led to the sustained increase in paraspeckle numbers in GBM cells, without affecting paraspeckle size (Fig. 3). Previous studies have shown NEAT1_1 is present in, but not essential for paraspeckle formation (27, 28, 44). However, whether deficiency of NEAT1_1 may affect the composition and function of paraspeckles, which are indicated in cellular stress responses, cell differentiation, and cancer progression, is not known (40, 41, 42). Interestingly, regardless of the essential function of paraspeckles in RNA processing (38, 78), many RNA processing pathways were downregulated in the NEAT1 ΔPAS GBM cell line which harbored elevated NEAT1_2 along with diminished NEAT1_1 (Fig. 6). These results raised a question as to whether NEAT1_1 is also important in governing paraspeckle integrity and function, which warrants further investigation in future studies.
So you get your cycler at the end of March? Well at least you have a day to look forward to now! Once you know your "prescription", you should ask about those smaller bags that Beach talks about in her initial posts before you get your initial cycler shipment. I didn't know they had cycler solution in less than 6000ml bags, 2 bags per box which are pretty darned heavy.
Hey, be kind to yourself during all this. I've made my share of mistakes. And once, my machine failed on the cassette test and I had to start over again, then another time, one of the solution bags had a bad kink in the "neck" and wouldn't allow the fluid to flow through properly, so had to put on a new bag. So yeah, "stuff" happens, both human and machine. Glad you had assistance going down your "checklist". I still follow mine every day so I don't forget something, though the day will come when I will forget, that's a given.
Importantly, loss of the QREs causes a significant increase of transcripts not cleaved at the NEAT1 PAS, which are detected by both primer pairs compared to the WT reporter (Fig. 5, C and D). This result clearly demonstrates that the QREs are critical for the efficient utilization of the NEAT1 PAS, which is essential for the biogenesis of NEAT1_1. These data suggest that mutating the QREs upstream of the NEAT1 PAS prevents QKI-5 binding, therefore ablating the effects of QKI-5 on modulating the usage of the NEAT1 PAS, which regulates NEAT1 isoforms.
To generate the NEAT1 PAS cleavage reporter, a region of the NEAT1 transcript spanning nucleotides 3235 to 4175 was amplified from BE(2)-M17 neuroblastoma cells via PCR followed by TA cloning using TOPO TA Cloning Kit. The plasmid was propagated in TOP10 Escherichia coli (Thermo Fisher, K450001SC) and the PCR insert was subcloned into a pEGFP-C2 vector using KpnI (Thermo Fisher Scientific, ER0521) and ApaI (Thermo Fisher Scientific, ER1411) restriction enzyme sites. The expected sequence in the reporter gene was confirmed by DNA sequencing of the plasmid. The predicted QREs upstream of the NEAT1 PAS were then mutated in the pEGFP-C2-NEAT1 cleavage reporter. Primers used for the first and second QRE mutations are provided in Table S3. Successful mutagenesis was confirmed by DNA sequencing of the plasmid.
NEAT1_2 is the architectural lncRNA indispensable for paraspeckle formation, whereas NEAT1_1 is not essential for paraspeckle formation (27, 28, 29, 36, 43, 44). Paraspeckles are phase-separated nonmembranous nuclear bodies present only in a subpopulation of mammalian cells (29, 32, 35, 36, 37, 38, 43, 44, 58). Of note, no previous studies have examined paraspeckles in human glioma cells. We sought to characterize paraspeckle formation in U373 GBM cells based on the colocalized NEAT1_2 fluorescence in situ hybridization (FISH) signal with the immunofluorescence (IF) of NONO, an essential RBP component of paraspeckles (35, 38, 59). As shown in Figure 3A, FISH signals of a NEAT1_2-specific probe largely colocalizes with NONO IF in U373 GBM cells, clearly indicating detection of paraspeckle foci (Fig. 3, representative paraspeckle foci are indicated by arrows). Importantly, both NEAT1 ΔPAS clones show marked increases of colocalized NEAT1_2 RNA-FISH signals with NONO IF compared to the parent control (Fig. 3A). Quantification of confocal images shows increased paraspeckle numbers (NEAT1 ΔPAS clone #1: 8.620 ± 1.000 and NEAT1 ΔPAS clone #2: 8.531 ± 1.083) compared to control (4.520 ± 0.479) (Fig. 3B). However, the area of each paraspeckle foci in confocal images is not significantly changed in either NEAT1 ΔPAS clone (0.792 ± 0.031 μm2 and 0.807 ± 0.031 μm2) compared to control (0.709 ± 0.031 μm2) (Fig. 3C). Furthermore, no change in NONO protein levels is observed by immunoblot, indicating that increased paraspeckle formation in NEAT1 ΔPAS clones is not due to increased expression of NONO protein (Fig. S3).
you can sleep in you don’t even have to close your transfer set unless you want to I wake up from drain pain then it goes to last fill so I’m awake usually so sometimes I will close it someday I have slept through it. The machince will say therapy has ended . I clean the transfer set clamp the bags and close transfer set . Then put in mini cap then unhook. You will have to write down what the machine say you can do it in morning too there three thing .
Four Monitor Mount | Height Adjustable | For Up to 32" Monitors, Mfg Code: ARMQUADSS.
Buy Whirlpool Control Board (Cb) Progr. online from BuySpares. Next working day UK delivery available or call 0844 557 5575.
RNA-FISH was conducted as previously described (96). Briefly, cells were grown on and fixed onto coverslips (Carolina, 633029) using 4% paraformaldehyde in 1× diethyl pyrocarbonate-phosphate buffered saline (DEPC-PBS) for 10 min at room temperature. Coverslips were washed with 1× DEPC-PBS at room temperature, followed by one wash with 2× saline sodium citrate (SSC) for 10 min at room temperature, and a final wash with prewarmed 2× SSC containing 10% formamide. Coverslips were incubated in prehybridization buffer for 1.5 h at 37 °C after which coverslips were incubated in hybridization buffer containing FISH probes (1:100) overnight at 37 °C. The next day, coverslips were washed with prewarmed 2× SSC containing 10% formamide, followed by additional washes with 2× SSC. Coverslips were incubated in blocking buffer for 1 h at room temperature. The coverslips were then incubated with a mouse mAb against NONO (35) at 1:500 dilution in blocking buffer for 1 h at room temperature. Coverslips were washed with 1× DEPC-PBS and then incubated with secondary anti-mouse Alexa Fluor 488 (Thermo Fisher Scientific, A11001) at 1:500 dilution and in blocking buffer for 1 h at room temperature. Coverslips were washed with 1× DEPC-PBS and mounted onto slides with ProLong Gold Antifade Mountant with 4′,6-diamidino-2-phenylindole (Life Technologies, P36935). The FISH probes used in this study include Human NEAT1 5′ Segment with Quasar 570 Dye (Stellaris, SMF-2036-1) and Human NEAT1 Middle Segment with Quasar 570 Dye (Stellaris, SMF-2037-1).
In this study, we provide the first evidence that NEAT1 isoforms are differentially dysregulated in patient-derived GBM GSCs. Moreover, we demonstrate that recognition and usage of the NEAT1 PAS is a crucial mechanism that regulates NEAT1 isoform biogenesis in glioma under control of the GBM risk factor QKI-5. A working model for the mechanism of QKI-5 in the regulation of NEAT1 isoforms is illustrated in Figure 8. Finally, we identified broad changes in the GBM transcriptome and biological pathways in glioma tumorigenesis that are caused by forced biogenesis of NEAT1_2 with reciprocal reduction of NEAT1_1 and provide the first evidence that NEAT1_2 is responsible for driving GBM cell migration.
DEGs with significantly changed splicing efficiency were identified using unpaired Student’s t test with p < 0.05 as a cut-off.
yes I have a cart I put the machine on and I have a plastic bin on wheels where we put the second bag my cart has a lower shelf but not a middle one and the nurse wanted the second bag in my plastic cart she said it to low on the bottom of my metal cart .With the tubing and since it on a wheel cart I can go all the way across the hall to my husband office where he sit in a lift chair most day if he not on his computer so I can check on him cause he a night owl he doesn’t go to sleep until 2 or 3 am.
Although extensive studies have explored the oncogenic roles of NEAT1, most studies focus on the function of NEAT1 in sponging specific miRNAs in the cytoplasm (33, 79). This is unlikely the function of NEAT1_2, which is restricted in the nuclei. Surprisingly, very few studies have addressed the impacts of NEAT1 on the human transcriptome. Only a limited number of studies have conducted RNA-seq and transcriptomic analysis upon manipulation of mouse Neat1, with fewer analyzing the impact of human NEAT1 (80, 81, 82, 83). To our knowledge, our studies provide the first characterization of GBM transcriptomic changes as a result of reciprocal alterations of NEAT1 isoform levels.
Yes bad drain pain still doctor said I may be more constipated than I think even if I go every day so today he wrote a prescription for lactulose solution we will see if that help. I did write in other post I past my accuracy test with a 2.2 this time .
ESP Plus · Quiet Units · Condenser-Air Cooled · Bohn · Larkin · Trenton · Condenser-Water Cooled · Coaxial · Mounting Bracket · Shell and Coil · Shell and Tube ...
I'm not sure about the eating effect. I know I don't always get my 2000 calories a day cause I have a hard time eating while dwelling and it seems like my whole day is taken up with exchanges (filling, dwelling, draining all day).
I hope my pain eases she did change the tidal today so we will see if that help. Will let y’all know when thought my my visit would be about 19 minutes but the social worker got hold of me and ask me to fill out these questionnaires a lot was how is kidney disease affecting you and question to see if you are depress.
An amount of 1 μg of total RNA from three biological replicates of U373 parent and NEAT1 ΔPAS clones (1 and 2) was used for poly-A-enriched RNA-seq library preparation using the TruSeq Stranded mRNA kit (Illumina, 20020594). Libraries were sequenced on an Illumina HiSeq platform (Admera Health, LLC) with a read length configuration of 150 paired end, targeting 80 M total reads per sample.
5 Oct 2012 — Full size C-Arms tend to be the most sought after model because with a full size, you can do just about any type of procedure. The open space in ...
the pd nurse order my first shipment she didn’t have them send me enough green cause my b/p was going so low she thought I would more yellow so my center is ordering more green o so they will take the charge cause they won’t be here before next order . My pd nurse told me to call them and tell then how many boxes of each I have left and Baxter will figure it out.
More than 98% of the human genome consists of noncoding sequences, which are extensively transcribed to produce noncoding RNAs (ncRNAs) (1, 2). Long noncoding RNAs (lncRNAs) are a family of ncRNAs that harbor greater than 200 nucleotides (nt) but lack protein-coding abilities (3, 4). Like mRNAs, lncRNAs are transcribed by RNA polymerase II, often undergo 5′ capping, splicing, and 3′ end polyadenylation (5, 6). lncRNAs are expressed in various cell types and tissues, which regulate broad gene networks through diverse mechanisms at the transcriptional and post-transcriptional levels (7, 8, 9, 10, 11). Functionally, lncRNAs influence essential biological processes including cell-cycle regulation, cell development, migration, and apoptosis (12, 13, 14). Interestingly, lncRNAs are much less conserved across evolution compared to protein coding genes (1, 15). Moreover, many lncRNAs are preferentially expressed in the brain and dysregulated in numerous neurodegenerative diseases (16, 17, 18, 19, 20, 21), as well as glioblastoma multiforme (GBM), the most common primary brain malignancy (22, 23, 24, 25).
When I had any beeping I just press stop and go and it would do what it was suppose to I only had beeping at the beginning . So that it.




 Neil
Neil 
 Neil
Neil